 |
 |
- Search
| Ann Pediatr Endocrinol Metab > Volume 24(1); 2019 > Article |
|
Abstract
Linear growth occurs at the growth plate. Therefore, genetic defects that interfere with the normal function of the growth plate can cause linear growth disorders. Many genetic causes of growth disorders have already been identified in humans. However, recent genome-wide approaches have broadened our knowledge of the mechanisms of linear growth, not only providing novel monogenic causes of growth disorders but also revealing single nucleotide polymorphisms in genes that affect height in the general population. The genes identified as causative of linear growth disorders are heterogeneous, playing a role in various growth-regulating mechanisms including those involving the extracellular matrix, intracellular signaling, paracrine signaling, endocrine signaling, and epigenetic regulation. Understanding the underlying genetic defects in linear growth is important for clinicians and researchers in order to provide proper diagnoses, management, and genetic counseling, as well as to develop better treatment approaches for children with growth disorders.
Height, as a measure of linear growth, is highly heritable, and genetics plays a major role in the regulation of linear growth [1]. To understand the genetic mechanisms of linear growth, various efforts have been made for decades, and recently genome-wide approaches such as genome-wide association (GWA) studies or exome/whole genome sequencing have been performed successfully, identifying many genomic loci associated with height variation in the general population as well as identifying monogenic changes that cause either syndromic or isolated short stature or overgrowth/tall stature [2-4]. Combining these newly discovered findings obtained from genome-wide approaches into important growth-regulating genes, we now possess a better understanding of human growth disorders [4]. It is now understood that: (1) linear growth occurs at the growth plate and many genes that affect height variation or cause growth disorders play major roles in growth plate biology [5]; (2) the type of mutation can determine the growth outcome; both loss or gain of function mutations can cause either short or tall stature. For example, loss-of-function mutations in NPR2 cause short stature while gain-of-function mutations in the same gene cause tall stature. On the other hand, loss-of-function mutations in FGFR3 cause tall stature while gain-of-function mutations cause short stature [4].; (3) the severity of the genetic abnormality can determine the severity of disease. For example, for some genes, a biallelic mutation can cause severe skeletal dysplasia while a monoallelic mutation in the same gene can cause isolated short stature, as seen in SHOX deficiency or ACAN mutations [6,7].; (4) lastly any genetic defect that alters the biology of chondrocytes in the growth plate can potentially cause growth disorders and these genetic defects can be very heterogeneous [4].
With recent remarkable advances in discoveries of genes responsible for regulating childhood growth disorders, it has become more important for pediatricians, pediatric endocrinologists and geneticists to understand the underlying genetic defects in children who present with growth concerns in order to provide genetic diagnoses, proper management, and genetic counseling. Therefore, this review aims to provide an update on genes known to cause human growth disorders specifically by focusing on those also found through GWA studies to be associated with normal height variation. The genes noted in this review are particularly important because single nucleotide polymorphisms (SNPs) in these genes (presumably causing mild changes) cause height variation in the normal range, but mutations (causing more deleterious changes) in the same gene cause distinctive linear growth disorders. Moreover, combined polymorphisms in these genes could potentially be genetic causes of isolated short stature in an oligo/polygenic manner. The listed genes reviewed here are categorized by their function in the growth plate with the mutations and their corresponding phenotype are described.
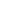
Linear growth occurs at the growth plate, a cartilaginous structure between the epiphysis and metaphysis (Fig. 1A). It is composed of 3 distinct layers; resting, proliferative and hypertrophic zones, which are temporally and spatially regulated (Fig. 1B) [8,9]. This regulation results in a unique differentiation process from resting chondrocytes to proliferating chondrocytes and finally hypertrophic chondrocytes. Ultimately, at the terminal zone of hypertrophic chondrocytes, hypertrophic chondrocytes undergo apoptosis and blood vessels invade, resulting in remodeling of the cartilage into newly formed bone [8]. Constant repetition of this process produces new bone and leads to the elongation of bones. Many genes play a role in this process and so far, hundreds of genes have been implicated in either human growth disorders or height variation by affecting chondrogenesis at the growth plate [5].
The growth mechanisms regulated by these genes can be divided into five different categories: (1) extracellular matrix, (2) intracellular signaling, (3) paracrine signaling, (4) endocrine signaling, and (5) epigenetic regulation (Fig. 1C).
Growth plate chondrocytes secrete an extracellular matrix that is essential for proper functioning of the growth plate. The extracellular matrix consists of collagens, proteoglycans, and glycosaminoglycans and along with secreted growth factors and the modifying enzymes, extracellular matrix serves as a dynamic scaffold that supports and orients growth plate chondrocyte differentiation [10]. Mutations in extracellular matrix-regulating genes can cause isolated short stature or severe skeletal dysplasia, supporting the idea that extracellular matrix plays a critical role in growth plate regulation in humans [11].
Aggrecan, encoded by ACAN, is a chondroitin sulfate proteoglycan, one of the main components of the extracellular matrix in growth plate cartilage that has been shown to modulate hedgehog signaling in a mouse model [12]. In aggrecan-null mice, growth plates were devoid of matrix, showed abnormal chondrocyte organization and differentiation, and the expression of important growth plate regulatory genes such as Col10a1, Sox9, and Ihh was also affected [13]. Therefore, alterations in the structure or function of aggrecan can significantly affect the growth plate cartilage resulting in a change in linear growth. Biallelic mutations in ACAN cause spondyloepimetaphyseal dysplasia and monoallelic mutations cause isolated short stature with or without advanced bone age causing early cessation of linear growth in most cases [6,14]. Patients with mutations in ACAN may also present with premature and severe osteoarthritis and osteochondritis dissecans [15,16]. Heterozygous ACAN mutations are one of common causes of idiopathic short stature; in a cohort of 428 patients with idiopathic short stature, 6 potential disease-causing heterozygous ACAN mutations were found (1.4% of patients) [17]. Thus, in children who have a family history of short stature and early-onset arthritis, ACAN mutations should be considered.
Fibrillins are the major components of microfibrils in the extracellular matrix, and 2 genes in particular, fibrillin 1 (FBN1) and fibrillin 2 (FBN2), have been implicated in growthregulating mechanisms, which regulate transforming growth factor beta (TGF-β) and bone morphogenetic factor (BMP) signaling in the extracellular matrix [3,18]. Monoallelic mutations in FBN1 give rise to acromelic dysplasia, a group of disorders that include Weill-Marchesani syndrome (WMS), acromicric dysplasia and gelophysic dysplasia [18].
Latent transforming growth factor-beta-binding protein 2 (LTBP2) is an extracellular matrix protein, which is a member of superfamily composed of three fibrillins and four other LTBP proteins and also participates in the regulation of TGF-β release in the extracellular matrix. Therefore biallelic mutations in LTBP2 can also cause WMS [19]. Patients with WMS present with short stature with brachydactyly, stiff joints, muscular build, thick skin, ectopia lentis and cardiac valvular problems [18,20]. Interestingly, mutations in FBN1 can also result in Marfan syndrome, an autosomal dominant inherited connective tissue disorder [21]. Individuals with Marfan syndrome have tall stature, skeletal and ocular abnormalities, and over 80% of Marfan syndrome patients have cardiovascular complications [22]. Although it has been proposed that mis-assembled microfibrils leads to decreased extracellular matrix integrity and the misregulation of TGFβ ligands leads to a Marfan syndrome phenotype, the precise mechanisms of how mutations in the same gene could cause 2 opposing phenotypes, short stature versus tall stature, is still unclear [23,24]. Monoallelic mutations in fibrillin 2 (FBN2) have been reported to cause a Marfanlike disorder with tall stature and congenital contractural arachnodactyly, which further confirms the importance of microfibrils in the regulation of linear growth [25]. However, it is not known yet if other mutations in FBN2 can also cause acromelic dysplasia.
Metalloproteinases are enzymes that degrade extracellular proteins. There are various kinds of metalloproteinases, such as matrix metalloproteinases (MMPs), a disintegrin and metalloproteinase (ADAMs), a disintegrin and metalloproteinase with thrombospondin motifs (ADALTSs) and ADAMTSlike proteins (ADAMTSLs). MMPs have been shown to play an integral role in regulating chondrocyte proliferation, extracellular matrix degradation, and release of extracellular matrix proteins [26]. However, there have not yet been reports of mutations in MMP-encoding genes causing skeletal defects in humans, although there has been a case report of Winchester syndrome due to a hypomorphic mutation in MMP14 [27]. The precise functions of these enzymes have not been wellestablished but many family members were identified by GWA studies to be associated with height variation (ADAM28, ADAMTS19, ADAMTS2, ADAMTS3, ADAMTS6, ADAMTSL1, ADAMTSL3) suggesting a possible role in linear growth. However, it is also possible that these genes may have been just innocent neighbors to other important genes associated with height variation. Among these, ADAMTS17 has been recognized as a modulator of fibrillin microfibrils, and therefore it is not surprising that biallelic mutations in ADAMTS17 cause Weill-Marchesani-like syndrome, where individuals present with short stature and spherophakia, but do not have brachydactyly and joint stiffness [28-30].
Cartilage fibrils are complex structures containing collagens II and XI with or without collagen IX and collagen XVI [31]. These, together with other fibrils containing small leucinerich proteins or proteoglycans, such as decorin, biglycan and fibromodulin, make up the fibrillar matrix [11]. The interaction of the fibrillar and extrafibrillar matrix then form a fibrillar periphery [31]. COL11A1 and COL27A1 are important for the organization of the proliferative zone, play a structural role in the pericellular extracellular matrix of the growth plate, and are thought to be important for the transition of cartilage to bone [32-34]. Mutations in these proteins underlie a broad spectrum of skeletal dysplasias (Table 1). Monoallelic mutations in COL2A1, COL11A1, COL11A2, COL9A1, COL9A2, and COL9A3 genes cause Stickler syndrome [35]. Heterozygous mutations in COL11A1 have also been implicated in Marshall syndrome, a chondrodysplasia that is phenotypically very similar to Stickler syndrome. There is some debate as to whether these disorders are separate entities or different expressions of the same disorder. They present with chondrodysplasias characterized by midfacial hypoplasia, myopia, sensorineural-hearing deficits, and short stature in the patient relative to their unaffected family members [36-38]. On the other hand, biallelic mutations cause fibrochondrogenesis 1, a skeletal dysplasia presenting with short limbs, flat midface and protrudent abdomen [39]. Furthermore, biallelic mutations in COL27A1 cause Steel syndrome [40-42]. The main clinical features of this disorder are bilateral hip and radial head dislocations, scoliosis, short stature, carpal coalitions, and pes cavus, with some patients also having facial dysmorphism [42]. Patients with monoallelic mutations in COL9A2 and MATN3 can present with multiple epiphyseal dysplasia, a clinically variable disease characterized by short stature and early onset osteoarthritis [43-45]. Conversely, biallelic mutations in COL9A2 and MATN3 cause a more severe form of skeletal dysplasia [44,47].
Various intracellular signaling pathways have been recognized as causative factors of linear growth impairment, and mutations in intracellular signaling have been identified to significantly impair linear growth. However, surprisingly, not many SNPs in these genes were identified to be associated with height variation. It may be because not all the functions of identified genes are well-established or the functions of these genes may be too critical and essential, such as transcriptional regulation, cyclic AMP (cAMP) production, and DNA repair, therefore preventing frequent changes (SNPs) that can affect linear growth.
SOX9 is a critical transcriptional factor for sex development during embryogenesis and for skeletal development and also plays an important role in chondrogenesis at the growth plate by regulating chondrocyte differentiation [48,49]. In humans, dominant mutations in SOX9 cause campomelic dysplasia (which means "bent limb" in Greek) with or without sex reversal [48,49].
The parathyroid hormone-related protein-Indian hedgehog (PTHrP-IHH) negative feedback loop is a major regulator of chondrogenesis in the growth plate, and PTHrP acts through the PTH1 receptor which activates both Gs and Gq family heterotrimeric G proteins [50]. The activation of Gs leads to cAMP production and protein kinase A (PKA) activation, and subsequent phosphorylation of the cAMP response elementbinding (CREB) family of transcription factors [51].
GNAS encodes the G protein alpha-subunit (Gs alpha), which, when activated, increases cAMP production to mediate downstream signaling [52]. Impairment of the cAMP signaling pathway through heterozygous loss-of-function mutations in Gs alpha leads to Albright's hereditary osteodystrophy (AHO) with or without hormone resistance. Maternal inheritance of Gs alpha mutations leads to AHO and pseudohypoparathyroidsim type 1a (PHP1a), which can include multihormone resistance, while patients with paternally inherited mutations of Gs alpha have only the AHO phenotype, termed pseudopseudohypo arathyroidism [52]. Phenotypically, patients with AHO present with short stature, brachydactyly, rounded face, and short neck [53]. Patients who have both AHO and PHP1a also have multihormone resistance to PTH, TSH, and gonadotropins, all of which act through G-protein-coupled receptors [52]. CREBBP encodes a nuclear transcriptional coactivator protein, CREB (cAMP-response element binding protein)-binding protein, that binds specifically to the PKA-phosphorylated form of the CREB protein. Rubinstein-Taybi syndrome (RSTS) is a rare condition characterized by short stature, intellectual disability, broad and deviated thumbs and halluces, and distinct craniofacial characteristics [54]. Over 65% of cases of RSTS are caused by dominant mutations in CREBBP [55].
PDE3A encodes a cyclic GMP (cGMP) and AMP (cAMP) phosphodiesterase 3A. Missense mutations in PDE3A have been shown to cause Mendelian hypertension and brachydactyly type E in seven unrelated families [56,57]. This syndrome has symptoms of short stature, brachydactyly, salt-independent and age-dependent hypertension, an increased fibroblast growth rate, and other cardiovascular complications. The identified mutations increase PKA-mediated PDE3A phosphorylation and increase cAMP-hydrolytic activity, thereby decreasing cAMP production [56].
Paracrine factors, such as fibroblast growth factors (FGFs), C-type natriuretic peptide (CNP), IHH, PTHrP encoded by PTHLH, and BMPs, play many critical roles in the growth plate, including chondrocyte proliferation, differentiation, hypertrophy, and matrix assembly [8]. Mutations in these genes have been identified to cause certain growth disorders with unique phenotype.
In the growth plate, IHH-PTHrP participates in a negative feedback loop that modulates chondrocyte differentiation [58]. Therefore, mutations in IHH, PTHLH, and the PTHrP receptor, PTH1R, result in disorders that affect skeletal growth. Mutations in IHH lead to brachydactyly type A1 and acrocapitofemoral dysplasia, and also can present with isolated short stature [59-61]. Mutations in PTHLH cause brachydactyly type E with short stature and the most prominent symptoms of these disorders are shortening of the metacarpals or metatarsals and short stature [62]. Loss-of-function mutations in PTH1R cause Blomstrand chondrodysplasia and gain-of-function mutations cause Jansen's metaphyseal chondrodysplasia, both of which are characterized primarily by short-limbed dwarfism [63,64].
The CNP is encoded by the NPPC gene and exerts its effects through the natriuretic peptide receptor B (NPR-B) encoded by NPR2 [65]. Loss of function mutations in NPPC and NPR2 cause short stature, while overexpression in NPPC or gain-of-function mutations in NPR2 cause overgrowth [66-69]. Binding of CNP to NPR-B stimulates guanylyl cyclase activity, increasing the synthesis of cGMP and activating the type 2 cGMP- dependent protein kinase, which then inhibits the MAPK pathway and antagonizes FGFR signaling [70,71]. Because FGFR signaling induces growth arrest of chondrocytes, CNP promotes chondrocyte proliferation. Biallelic NPR2 mutations cause acromesomelic dysplasia, Maroteaux type (AMDM), an autosomal recessive skeletal dysplasia characterized by severe short stature and short limbs [72]. Multiple mutations causing loss of function in NPR-B have been associated with AMDM. Nonsense mutations in the intracellular kinase homology domain (KHD) of NPR-B lead to impaired cGMP production, providing evidence that this domain is essential for skeletal growth [73,74]. Gain-of-function mutations in the NPR2 gene have also been described, and these patients have tall stature and macrodactyly of the big toes [69,74,75]. It has been shown that a gain-of-function mutation in the KHD domain of NPR-B results in an increase in cGMP activity compared to the wildtype protein [76].
The GDF5 gene encodes growth differentiation factor 5 (also known as cartilage-derived morphogenic protein) which acts through BMP receptors. Mutations in this gene can cause brachydactyly (type A1, A2, and C) with short stature, chondrodysplasia - Grebe type, characterized by severe dwarfism, disproportionate short stature, and brachydactyly, and Hunter-Thompson type, characterized by skeletal abnormalities that are restricted to the limbs and limb joints [77-81]. Early studies showed that GDF5 is primarily expressed in long bones during human embryonic development [82] and recent studies revealed that GDF5 enhances cartilage formation and GDF5 and TGFβ synergistically induce mesenchymal stem cell differentiation into chondrocytes, providing evidence for its important role in cartilage development [83,84].
The canonical WNT signaling pathway is important for cell proliferation and cell fate change during development. Studies in Wnt5a-null mice showed a defect in cell polarization in the growth plate [85]. Dominant mutations in WNT5A are associated with Robinow syndrome, a disorder characterized by short stature, limb shortening, genital hypoplasia, and craniofacial abnormalities [86,87].
IGF2 encodes insulin-like growth factor 2, an important fetal growth factor that binds to the type 1 IGF receptor to regulate cell proliferation. Recently, studies showed that growth plates in Igf2- null mice show a disproportionally larger hypertrophic zone and delayed secondary ossification center, and, in humans, paternally inherited mutations in IGF2 in human cause postnatal growth failure in addition to prenatal growth [88,89]. The dysmorphic features of affected family members were similar to those of individuals with IGF-2 deficiency seen in Silver-Russell syndrome [89]. IGFBP2BP2 and IGF2BP3, encoding the IGF2 binding proteins, are identified to be associated with height variation but mutations have not yet been found in human disorders.
Endocrine signaling pathways are the clinical causes most extensively evaluated in children who present with short stature since endocrine causes and hormonal abnormalities are relatively easily tested in clinical practice. Growth hormone, thyroid hormone, glucocorticoids, and sex steroids affect chondrocyte proliferation and hypertrophy in the growth plate, altering linear growth if there is a defect in the corresponding axis.
The GH-IGF1 axis is one of the main regulatory systems that controls chondrogenesis in the growth plate, whereby GH acts directly on the growth plate to stimulate chondrogenesis or indirectly through circulating and local insulin-like growth factor-1 (IGF-1) [90,91]. There have been multiple genes identified that cause growth hormone deficiency or impair downstream signaling of the GH-IGF1 axis. For the scope of this review, only genes identified by GWA studies are mentioned. Mutations in the IGF-1 receptor gene, IGF1R, can cause both prenatal and postnatal growth failure. Monoallelic mutations may cause short stature in children born small for gestational age [92]. Biallelic mutations cause severe short stature [93]. Fibroblast cultures from these patients showed decreased IGF1R function and a reduced number of IGF1 receptors, which could lead to IGF1 insensitivity [93]. SNPs in IGFBP3, IGFBP4, STC2, PAPPA, PAPPA2, which regulate the free form of IGF-1 by functioning as a binding protein (IGFBP3, IGFBP4), binding-protein-cleaving enzymes (PAPPA, PAPPA2), or as a regulator of PAPPA (STC2), have been identified by GWA studies to be associated with height variation, likely by affecting IGF-1 bioavailability and therefore the GH-IGF1 axis. However, only mutations in PAPPA2, encoding Pappalysin-2, a protein that cleaves IGFBP-3 and -5, have been identified in humans where the patients have decreased free IGF-1 levels [94].
PIK3R1 is a gene involved in insulin signaling. Patients with mutations in PIK3R1 have SHORT syndrome, hallmarks of which include short stature, hyperextensibility of joints, ocular depression, Rieger anomaly, teething decay, and insulin resistance [95-97]. However, the role of PIK3R1 in the growth plate is not yet known.
Estrogen plays an important role in skeletal maturation and mineralization, and defects in estrogen signaling can lead to a variety of disorders that manifest with short or tall stature by altering the timing of growth plate senescence [98]. Since estrogen normally accelerates growth plate senescence and skeletal maturation, children with aromatase deficiency, a rare autosomal recessive disorder caused by mutations in the CYP19A1 gene, present with delayed bone age and delayed puberty which lead to tall stature. This presentation is due to a lack of estrogen converted from androgen [99,100]. On the other hand, overexpression of CYP19A1 can result in aromatase excess syndrome. Affected adults present with short stature, gynecomastia in males, and macromastia in females [101]. Patients with mutations in the estrogen receptor gene ESR1 have estrogen resistance. To date, there have been 5 patients described who have mutations in ESR1, with varying phenotypes and varying degrees of estrogen insensitivity, likely due to different mutation sites and differences in penetrance of the particular mutation [102,103]. However, all patients had delayed skeletal maturation, with some having tall stature in adulthood also due to a lack of estrogen effects on the growth plate [104].
DNA and histone methylation are one of the main gene regulatory mechanisms and play crucial roles in growth and development. The best example of epigenetic regulation of growth is the epimutations in the telomeric imprinting control region 1 (ICR1), which alters H19 and IGF2 expression. Hypermethylation of the ICR1 causes biallelic expression of IGF2 presenting as Beckwith-Wiedemann syndrome (BWS) (overgrowth) while hypomethylation of the ICR1 causes suppression of IGF2 expression presenting as Silver-Russell syndrome (short stature) [105]. Moreover, many genes involved in DNA methylation have been identified to cause growth disorders, especially overgrowth syndromes. Studies have found de novo mutations in DNMT3A, a gene that encodes a DNA methyltransferase, which establishes methylation during embryogenesis [106]. These mutations cause an overgrowth syndrome called DNMT3A syndrome. Patients present with tall stature, intellectual disability, and distinctive facial appearance with a round face, heavy, horizontal eyebrows, and narrow palpebral fissures [106,107]. In addition to DNA methylation, histone methylation has been implicated in growth disorders. HIST1H1E encodes histone H1.4, and in humans, H1.4 mediates the formation of higher-order chromatin structures, thus regulating the accessibility of histone modification factors, regulatory proteins, and chromatin modifiers to target sites on this chromatin structure [108]. Mutations in this gene cause Rahman syndrome. Individuals with this disease have overgrowth with varying degrees of intellectual disability and a distinctive facial appearance in childhood - full cheeks, high hairlines, and telecanthus [108]. Additionally, monoallelic mutations in genes that encode other histone methyltransferases cause Weaver syndrome and Sotos syndrome. Weaver syndrome is caused by mutations in EZH2 (Enhancer of Zeste homolog 2) [109,110]. This gene encodes a histone methyltransferase that is part of the polycomb repressive complex 2, which catalyzes the methylation of lysine residue 27 of histone 3 (H3K27), thus causing transcriptional repression [111]. Patients with Weaver syndrome have tall stature, advanced bone age, dysmorphic facial features, and variable learning disability [109,110]. Sotos syndrome is another overgrowth syndrome characterized by overgrowth, advanced bone age, dysmorphic facial features, and intellectual disability, and patients with Sotos syndrome also often have brain anomalies and seizures [112]. However, in contrast to Weaver syndrome, Sotos syndrome and Sotoslike syndrome are caused by mutations in genes that cause transcriptional activation: NSD1 and SETD2 [112,113]. These 2 genes encode histone methyltransferases which catalyze the methylation of histone 3 lysine 36 to allow for transcriptional activation [112,113]. Interestingly, mutations in NSD1 have been detected in 2 patients with BWS, another overgrowth condition characterized by macroglossia, abdominal wall defects, visceromegaly, embryonic tumors, hemihyperplasia, and neonatal hypoglycemia [114].
In this review, we reviewed the genes that cause linear growth disorders in humans, focusing on their functions at the growth plate and their clinical phenotype. These genes are also identified by GWA studies which suggest that a mild change of function or expression in these genes may modulate linear growth within the normal ranges. The genetic evidence supports the idea that genes causing linear growth are highly heterogeneous, so that any regulatory mechanisms that alter chondrogenesis in the growth plate could be the genetic cause of growth disorders, either syndromic or isolated. With the aid of genome-wide approaches, it is expected that additional regulating mechanisms in growth disorders will be discovered and broaden our knowledge of the underlying causes of these disorders.
ACKNOWLEDGMENTS
This work was supported by the Intramural Research Program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development, NIH (ZIA HD000640).
Fig. 1.
Mechanisms regulating chondrocytes in the growth plate. (A) Growth plate on X-ray. Black box indicates where growth plate is located. (B) Diagram of the 3 layers in the growth plate. (C) Diagram of chondrocyte-regulating mechanisms. RZ, resting zone; PZ, proliferative zone; HZ, hypertrophic zone. Modified from Jee YH, et al. Endocrinol Metab Clin North Am 2017;46:259-81 [15].
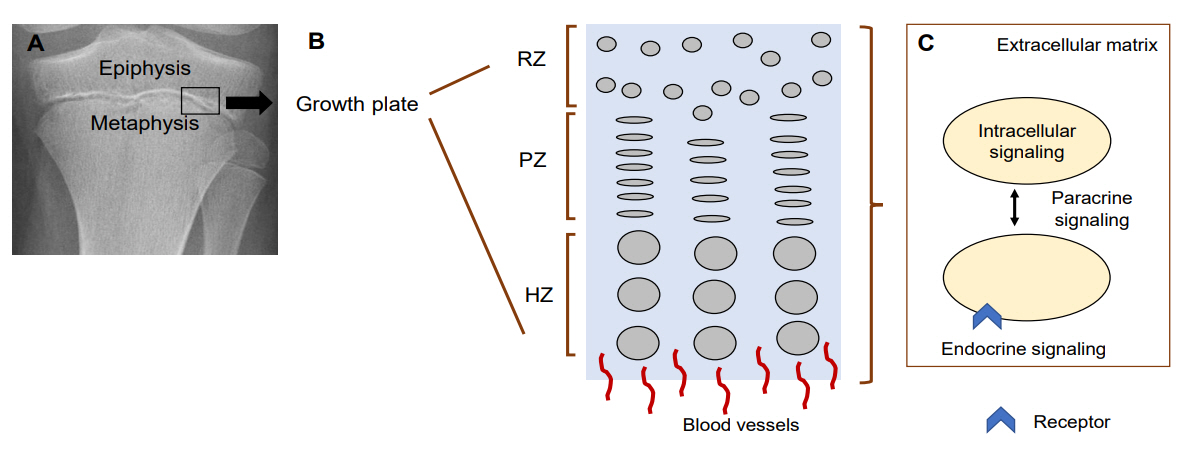
Table 1.
Genetic causes of linear growth disorders
| Gene | Protein | Disorders | Clinical presentation | Height-associated SNP ID [2,3] |
|---|---|---|---|---|
| Extracellular matrix | ||||
|
|
||||
| ACAN | Aggrecan | Isolated short stature (monoallelic) | Short stature with or without advanced bone age, short legs, long arms, midface hypoplasia, osteochondritis dissecans [14-16] | rs2280470, rs3817428, rs11633371, rs16942341 |
| Spondyloepimetaphyseal dysplasia (biallelic) | Extreme short stature, increased upper/lower segment ratio, rhizomelia, mesomelia, brachydactyly, midface hypoplasia, short neck, lumbar lordosis, early onset arthritis [6,15] | |||
| ADAMTS17 | ADAM metallopeptidase with thrombospondin type 1 Motif 17 | Weill-Marchesani syndrome (biallelic) | Short stature without brachydactyly and joint stiffness, short hands and feet, spherophakia, myopia, cataract, thick skin [27-29] | rs2573625, rs4246302, rs4548838 |
| COL9A2 | Collagen type IX alpha 2 chain | Multiple epiphyseal dysplasia (monoallelic) Stickler syndrome (biallelic) | Short stature, early onset osteoarthritis, brachydactyly [41,42] | rs209918 |
| COL11A1 | Collagen type XI alpha 1 chain | Stickler syndrome, Marshall syndrome (monoallelic) | Short stature, midface hypoplasia, short nose, myopia, sensorineural-hearing deficits [34-36] | rs7517682 |
| Fibrochondrogenesis 1 (biallelic) | Short-limbed skeletal dysplasia, flat midface, small nose with anteverted nares, relatively normal hands and feet, small thorax [37] | |||
| COL27A1 | Collagen type XXVII alpha 1 chain | Steel syndrome (biallelic) | Short stature, bilateral hip and radial head dislocations, scoliosis, carpal coalitions, pes cavus, facial dysmorphism (long oval face, prominent forehead, hypertelorism, broad nasal bridge) [38-40] | rs999599 |
| FBN1 | Fibrillin 1 | Acromelic dysplasia;Weill-Marchesani syndrome, acromicric dysplasia, gelophysic dysplasia (monoallelic) | Short stature, spherophakia, brachydactyly, joint stiffness, thick skin, cardiac valvular abnormalities [17] | rs1036477 |
| Marfan syndrome (monoallelic) | Tall stature, arachnodactyly, dolichostenomelia, joint hypermobility, ectopia lentis, aortic root enlargement, aortic dissection [20-23] | |||
| FBN2 | Fibrillin 2 | Marfan like-disorder (monoallelic) | Tall stature, congenital contractural arachnodactyly, dolichostenomelia, joint hypermobility, without occular or cardiac manifestations [24] | rs26024 |
| LTBP2 | Latent transforming growth factor beta binding protein 2 | Weill-Marchesani syndrome (biallelic) | Short stature, brachydactyly, joint stiffness, short hands and feet, spherophakia, myopia, cataract, thick skin [18] | rs862034, rs10140101 |
| MATN3 | Matrilin-3 | Multiple epiphyseal dysplasia (monoallelic) | Short stature, early onset osteoarthritis, brachydactyly [43] disproportionate early-onset dwarfism, bowing of lower limbs, lumbar lordosis, and normal hands, abnormal skeletal X-rays (short, wide, and stocky long bones with severe epiphyseal and metaphyseal changes, hypoplastic iliac bones, and flat-ovoid vertebral bodies) [45] | rs52826764 |
| Spondyloepimetaphyseal dysplasia (biallelic) | ||||
|
|
||||
| Intracellular signaling | ||||
|
|
||||
| CREBBP | CREB (cAMP response elementbinding protein) binding protein | Rubinstein-Taybi syndrome (monoallelic) | Short stature, intellectual disability, broad and deviated thumbs and halluces, distinct craniofacial characteristics [52,53] | rs129963 |
| GNAS | Guanine nucleotide binding protein, alpha stimulating subunit | Albright's hereditary osteodystrophy with hormone resistance-pseudohypoparathyroidism (monoallelic, maternally inherited) | Short stature, brachydactyly, rounded face, short neck, centripetal obesity, developmental delay, subcutaneous ossifications, multihormonal resistance to PTH, TSH and gonadotropins [50,51] | rs2057291 |
| Albright's hereditary osteodystrophy without hormone resistance-pseudohypoparathyroidism (monoallelic, paternally inherited) | Short stature, brachydactyly, rounded face, short neck, subcutaneous ossification [50,51] | |||
| PDE3A | Phosphodiesterase 3A | Mendelian hypertension with brachydactyly type E (monoallelic) | Short stature, brachydactyly, salt-independent and age-dependent hypertension, an increased fibroblast growth rate, and other cardiovascular complications [54,55] | rs4326884 |
| SOX9 | SRY-box 9 | Campomelic dysplasia (monoallelic) | Congenital bowing of long bones, with or without sex reversal [46,47] | rs10083886 |
|
|
||||
| Paracrine signaling | ||||
|
|
||||
| GDF5 | Growth differentiation factor 5 (cartilage-derived morphogenic protein 1) | Brachydactyly, type A1, A2, C (monoallelic) | Short stature and brachydactyly [75-77] | rs143384 |
| Chondrodysplasia - Grebe type (biallelic) | Severe dwarfism with micromelia, deformation of limbs, disproportionate short stature, brachydactyly [77,78] | |||
| Chondrodysplasia - Hunter - Thompson type (biallelic) | Short long bones (most severe in hands and feet), joint dislocations, normal craniofacial skeleton [79] | |||
| IGF2 | Insulin-like growth factor II | Growth restriction (monoallelic) | Paternally inherited, pre- and postnatal growth restriction with the features of Silver-Russell syndrome [87] | rs4320932 |
| IHH | Indian hedgehog | Brachydactyly type A1 (monoallelic) | Shortening of the metacarpals or metatarsals, and short stature [57,58] | rs142036701 |
| Acrocapitofemoral dysplasia (biallelic) | Short stature with short limbs, shortening of the metacarpals or metatarsals, thorax deformities, cone-shaped epiphyses in hands and femur head [59] | |||
| NPPC | C-type natriuretic peptide | Isolated short stature (monoallelic, loss-of-function mutation) | Short stature [64] | rs749052 |
| Overgrowth (NPPC overexpression) | Tall stature, long halluses, spine and joint deformities, Marfanoid habitus [66] | |||
| NPR2 | Natriuretic peptide receptor B | Isolated short stature (monoallelic) | Isolated short stature [65] | rs3763631 |
| Acromesomelic dysplasia, Maroteaux type (biallelic) | Severe short stature, short bowing limbs, phalangeal-metacarpal abnormalities, brachydactyly of hands and feet [70] | |||
| Overgrowth (gain-of-function mutation, monoallelic) | Tall stature, macrodactyly of big toes [67] | |||
| PTH1R | PTH/PTH-related peptide receptor, type 1 | Blomstrand chondrodysplasia (loss-of-function mutation, biallelic) | Extremely advanced endochondral bone maturation, increased bone density, [61] | rsl21434601 |
| Jansen's metaphyseal chondrodysplasia (gain-of-function mutation, monoallelic) | Short-limbed dwarfism, hypercalcemia and hypophosphatemia, normal or undetectable PTH and PTHrP [62] | |||
| PTHLH | Parathyroid hormone-related peptide | Brachydactyly type E (monoallelic) | Short stature, shortening of the metacarpals or metatarsals [60] | rs10492364 |
| WNT5A | Wnt family member 5A | Robinow syndrome (monoallelic) | Short stature, limb shortening, genital hypoplasia, mandibular hypoplasia, hypertelorism [84,85] | rs2034172 |
|
|
||||
| Endocrine signaling | ||||
|
|
||||
| CYP19A1 | Aromatase (cytochrome p450 family 19 subfamily A member 1) | Aromatase excess syndrome (monoallelic) | Short stature, gynecomastia in males, macromastia in females [98] | rs16964211 |
| Aromatase deficiency (biallelic) | Tall stature compared to midparental height, delayed bone age, delayed puberty, ambiguous genitalia at birth, elevated androgens, low estrogen [96,97] | |||
| ESR1 | Estrogen receptor | Estrogen resistance (biallelic) | Tall stature (variable), estrogen insensitivity, delayed skeletal maturation, no breast development in women [99,100] | rs3020418, rs6902771 |
| IGF1R | Insulin-like growth factor I receptor | Isolated short stature (monoallelic) | Small for gestational age, short stature [90] | rs2871865 |
| Prenatal and postnatal growth failure (biallelic) | Intrauterine growth retardation, severe postnatal growth failure, microcephaly, developmental delay [91] | |||
| PIK3R1 | Phosphoinositide-3-kinase regulatory subunit 1 | SHORT syndrome (monoallelic) | Short stature, hyperextensibility of joints, ocular defect (Rieger anomaly), lipodystrophy, insulin resistance, low birth weight, delayed bone age, triangular facies, hypoplastic nasal alae, low set ears [93-95] | rs9291926 |
|
|
||||
| Epigenetic regulation | ||||
|
|
||||
| DNMT3A | DNA methyltransferase 3 alpha | DNMT3A syndrome (monoallelic) | Tall stature, intellectual disability, round face, heavy and horizontal eyebrows, narrow palpebral fissures [103,104] | rs2289195, rs10460566 |
| EZH2 | Enhancer of Zeste homolog 2 | Weaver syndrome (monoallelic) | Tall stature, advanced bone age, macrocephaly, hypertelorism, retrognathia, variable learning disability [106,107] | rs822531 |
| HIST1H1E | H1 histone family, member 4 | Rahman syndrome (monoallelic) | Variable stature, intellectual disability and head circumference, facial dysmorphism (full cheeks, high hairlines, telecanthus) [105] | rs4141885 |
| NSD1 | Nuclear receptor binding SET domain protein 1 | Sotos syndrome (monoallelic) | Overgrowth, advanced bone age, macrocephaly, pointed chin, receding hairline, downslanting palpebral fissures, intellectual disability, brain anomalies, seizures [109] | rs11950938, rs12055154 |
| Beckwith-Weidemann syndrome (monoallelic) | Overgrowth, ear and renal anomalies, macroglossia, abdominal wall defects, visceromegaly, embryonic tumors, hemihyperplasia, neonatal hypoglycemia [111] | |||
| SETD2 | SET domain containing 2 | Sotos syndrome (monoallelic) | Overgrowth, macrocephaly, finical dysmorphism, long and large hands and feet, advanced bone age, mild intellectual disability [110] | rs76208147 |
References
1. Lango Allen H, Estrada K, Lettre G, Berndt SI, Weedon MN, Rivadeneira F, et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010;467:832–8.




2. Chan Y, Salem RM, Hsu YH, McMahon G, Pers TH, Vedantam S, et al. Genome-wide analysis of body proportion classifies height-associated variants by mechanism of action and implicates genes important for skeletal development. Am J Hum Genet 2015;96:695–708.



3. Marouli E, Graff M, Medina-Gomez C, Lo KS, Wood AR, Kjaer TR, et al. Rare and low-frequency coding variants alter human adult height. Nature 2017;542:186–90.




4. Baron J, Sävendahl L, De Luca F, Dauber A, Phillip M, Wit JM, et al. Short and tall stature: a new paradigm emerges. Nat Rev Endocrinol 2015;11:735–46.




5. Lui JC, Nilsson O, Chan Y, Palmer CD, Andrade AC, Hirschhorn JN, et al. Synthesizing genome-wide association studies and expression microarray reveals novel genes that act in the human growth plate to modulate height. Hum Mol Genet 2012;21:5193–201.




6. Tompson SW, Merriman B, Funari VA, Fresquet M, Lachman RS, Rimoin DL, et al. A recessive skeletal dysplasia, SEMD aggrecan type, results from a missense mutation affecting the C-type lectin domain of aggrecan. Am J Hum Genet 2009;84:72–9.



7. Nilsson O, Guo MH, Dunbar N, Popovic J, Flynn D, Jacobsen C, et al. Short stature, accelerated bone maturation, and early growth cessation due to heterozygous aggrecan mutations. J Clin Endocrinol Metab 2014;99:E1510–8.




9. Lui JC, Andrade AC, Forcinito P, Hegde A, Chen W, Baron J, et al. Spatial and temporal regulation of gene expression in the mammalian growth plate. Bone 2010;46:1380–90.



10. Melrose J, Shu C, Whitelock JM, Lord MS. The cartilage extracellular matrix as a transient developmental scaffold for growth plate maturation. Matrix Biol 2016;52-54:363–83.


11. Myllyharju J. Extracellular matrix and developing growth plate. Curr Osteoporos Rep 2014;12:439–45.



12. Cortes M, Baria AT, Schwartz NB. Sulfation of chondroitin sulfate proteoglycans is necessary for proper Indian hedgehog signaling in the developing growth plate. Development 2009;136:1697–706.



13. Lauing KL, Cortes M, Domowicz MS, Henry JG, Baria AT, Schwartz NB. Aggrecan is required for growth plate cytoarchitecture and differentiation. Dev Biol 2014;396:224–36.



14. Tatsi C, Gkourogianni A, Mohnike K, DeArment D, Witchel S, Andrade AC, et al. Aggrecan mutations in nonfamilial short stature and short stature without accelerated skeletal maturation. J Endocr Soc 2017;1:1006–11.




15. Gleghorn L, Ramesar R, Beighton P, Wallis G. A mutation in the variable repeat region of the aggrecan gene (AGC1) causes a form of spondyloepiphyseal dysplasia associated with severe, premature osteoarthritis. Am J Hum Genet 2005;77:484–90.



16. Stattin EL, Wiklund F, Lindblom K, Onnerfjord P, Jonsson BA, Tegner Y, et al. A missense mutation in the aggrecan C-type lectin domain disrupts extracellular matrix interactions and causes dominant familial osteochondritis dissecans. Am J Hum Genet 2010;86:126–37.



17. Hauer NN, Sticht H, Boppudi S, Büttner C, Kraus C, Trautmann U, et al. Genetic screening confirms heterozygous mutations in ACAN as a major cause of idiopathic short stature. Sci Rep 2017;7:12225.



18. Sakai LY, Keene DR. Fibrillin protein pleiotropy: Acromelic dysplasias. Matrix Biol 2018;Sep 13 [Epub]. pii: S0945-053X(18)30312-3. https://doi.org/10.1016/j.matbio.2018.09.005.

19. Haji-Seyed-Javadi R, Jelodari-Mamaghani S, Paylakhi SH, Yazdani S, Nilforushan N, Fan JB, et al. LTBP2 mutations cause Weill-Marchesani and Weill-Marchesani-like syndrome and affect disruptions in the extracellular matrix. Hum Mutat 2012;33:1182–7.


20. Kojuri J, Razeghinejad MR, Aslani A. Cardiac findings in Weill-Marchesani syndrome. Am J Med Genet A 2007;143A:2062–4.


21. Grau U, Klein HG, Detter C, Mair H, Welz A, Seidel D, et al. A novel mutation in the neonatal region of the fibrillin (FBN)1 gene associated with a classical phenotype of Marfan syndrome (MfS). Mutations in brief no. 163. Online. Hum Mutat 1998;12:137.

22. Cook JR, Ramirez F. Clinical, diagnostic, and therapeutic aspects of the Marfan syndrome. Adv Exp Med Biol 2014;802:77–94.


23. Park JW, Yan L, Stoddard C, Wang X, Yue Z, Crandall L, et al. Recapitulating and correcting Marfan syndrome in a cellular model. Int J Biol Sci 2017;13:588–603.



24. Cecchi A, Ogawa N, Martinez HR, Carlson A, Fan Y, Penny DJ, et al. Missense mutations in FBN1 exons 41 and 42 cause Weill-Marchesani syndrome with thoracic aortic disease and Marfan syndrome. Am J Med Genet A 2013;161A:2305–10.


25. Putnam EA, Zhang H, Ramirez F, Milewicz DM. Fibrillin-2 (FBN2) mutations result in the Marfan-like disorder, congenital contractural arachnodactyly. Nat Genet 1995;11:456–8.



26. Malemud CJ. Matrix metalloproteinases: role in skeletal development and growth plate disorders. Front Biosci 2006;11:1702–15.


27. de Vos IJ, Tao EY, Ong SL, Goggi JL, Scerri T, Wilson GR, et al. Functional analysis of a hypomorphic allele shows that MMP14 catalytic activity is the prime determinant of the Winchester syndrome phenotype. Hum Mol Genet 2018;27:2775–88.




28. Shah MH, Bhat V, Shetty JS, Kumar A. Whole exome sequencing identifies a novel splice-site mutation in ADAMTS17 in an Indian family with Weill-Marchesani syndrome. Mol Vis 2014;20:790–6.


29. Morales J, Al-Sharif L, Khalil DS, Shinwari JM, Bavi P, Al-Mahrouqi RA, et al. Homozygous mutations in ADAMTS10 and ADAMTS17 cause lenticular myopia, ectopia lentis, glaucoma, spherophakia, and short stature. Am J Hum Genet 2009;85:558–68.



30. Hubmacher D, Apte SS. ADAMTS proteins as modulators of microfibril formation and function. Matrix Biol 2015;47:34–43.



31. Zaucke F, Grässel S. Genetic mouse models for the functional analysis of the perifibrillar components collagen IX, COMP and matrilin-3: Implications for growth cartilage differentiation and endochondral ossification. Histol Histopathol 2009;24:1067–79.

32. Plumb DA, Ferrara L, Torbica T, Knowles L, Mironov A Jr, Kadler KE, et al. Collagen XXVII organises the pericellular matrix in the growth plate. PLoS One 2011;6:e29422.



33. Hjorten R, Hansen U, Underwood RA, Telfer HE, Fernandes RJ, Krakow D, et al. Type XXVII collagen at the transition of cartilage to bone during skeletogenesis. Bone 2007;41:535–42.



34. Li Y, Lacerda DA, Warman ML, Beier DR, Yoshioka H, Ninomiya Y, et al. A fibrillar collagen gene, Col11a1, is essential for skeletal morphogenesis. Cell 1995;80:423–30.


35. Van Camp G, Snoeckx RL, Hilgert N, van den Ende J, Fukuoka H, Wagatsuma M, et al. A new autosomal recessive form of Stickler syndrome is caused by a mutation in the COL9A1 gene. Am J Hum Genet 2006;79:449–57.



36. Majava M, Hoornaert KP, Bartholdi D, Bouma MC, Bouman K, Carrera M, et al. A report on 10 new patients with heterozygous mutations in the COL11A1 gene and a review of genotype-phenotype correlations in type XI collagenopathies. Am J Med Genet A 2007;143A:258–64.


37. Annunen S, Körkkö J, Czarny M, Warman ML, Brunner HG, Kääriäinen H, et al. Splicing mutations of 54-bp exons in the COL11A1 gene cause Marshall syndrome, but other mutations cause overlapping Marshall/Stickler phenotypes. Am J Hum Genet 1999;65:974–83.



38. Griffith AJ, Sprunger LK, Sirko-Osadsa DA, Tiller GE, Meisler MH, Warman ML. Marshall syndrome associated with a splicing defect at the COL11A1 locus. Am J Hum Genet 1998;62:816–23.



39. Tompson SW, Bacino CA, Safina NP, Bober MB, Proud VK, Funari T, et al. Fibrochondrogenesis results from mutations in the COL11A1 type XI collagen gene. Am J Hum Genet 2010;87:708–12.



40. Gariballa N, Ben-Mahmoud A, Komara M, Al-Shamsi AM, John A, Ali BR, et al. A novel aberrant splice site mutation in COL27A1 is responsible for Steel syndrome and extension of the phenotype to include hearing loss. Am J Med Genet A 2017;173:1257–63.


41. Kotabagi S, Shah H, Shukla A, Girisha KM. Second family provides further evidence for causation of Steel syndrome by biallelic mutations in COL27A1. Clin Genet 2017;92:323–6.


42. Gonzaga-Jauregui C, Gamble CN, Yuan B, Penney S, Jhangiani S, Muzny DM, et al. Mutations in COL27A1 cause Steel syndrome and suggest a founder mutation effect in the Puerto Rican population. Eur J Hum Genet 2015;23:342–6.



43. Jackson GC, Marcus-Soekarman D, Stolte-Dijkstra I, Verrips A, Taylor JA, Briggs MD. Type IX collagen gene mutations can result in multiple epiphyseal dysplasia that is associated with osteochondritis dissecans and a mild myopathy. Am J Med Genet A 2010;152A:863–9.



44. Baker S, Booth C, Fillman C, Shapiro M, Blair MP, Hyland JC, et al. A loss of function mutation in the COL9A2 gene causes autosomal recessive Stickler syndrome. Am J Med Genet A 2011;155A:1668–72.


45. Mäkitie O, Mortier GR, Czarny-Ratajczak M, Wright MJ, Suri M, Rogala P, et al. Clinical and radiographic findings in multiple epiphyseal dysplasia caused by MATN3 mutations: description of 12 patients. Am J Med Genet A 2004;125A:278–84.


46. Baker S, Booth C, Fillman C, Shapiro M, Blair MP, Hyland JC, et al. A loss of function mutation in the COL9A2 gene causes autosomal recessive Stickler syndrome. Am J Med Genet A 2011;155A:1668–72.


47. Borochowitz ZU, Scheffer D, Adir V, Dagoneau N, Munnich A, Cormier-Daire V. Spondylo-epi-metaphyseal dysplasia (SEMD) matrilin 3 type: homozygote matrilin 3 mutation in a novel form of SEMD. J Med Genet 2004;41:366–72.



48. Wagner T, Wirth J, Meyer J, Zabel B, Held M, Zimmer J, et al. Autosomal sex reversal and campomelic dysplasia are caused by mutations in and around the SRY-related gene SOX9. Cell 1994;79:1111–20.


49. Foster JW, Dominguez-Steglich MA, Guioli S, Kwok C, Weller PA, Stevanović M, et al. Campomelic dysplasia and autosomal sex reversal caused by mutations in an SRYrelated gene. Nature 1994;372:525–30.



50. Abou-Samra AB, Jüppner H, Force T, Freeman MW, Kong XF, Schipani E, et al. Expression cloning of a common receptor for parathyroid hormone and parathyroid hormone-related peptide from rat osteoblast-like cells: a single receptor stimulates intracellular accumulation of both cAMP and inositol trisphosphates and increases intracellular free calcium. Proc Natl Acad Sci U S A 1992;89:2732–6.



51. Singh AT, Gilchrist A, Voyno-Yasenetskaya T, Radeff-Huang JM, Stern PH. G alpha12/G alpha13 subunits of heterotrimeric G proteins mediate parathyroid hormone activation of phospholipase D in UMR-106 osteoblastic cells. Endocrinology 2005;146:2171–5.



53. Mantovani G, Spada A. Mutations in the Gs alpha gene causing hormone resistance. Best Pract Res Clin Endocrinol Metab 2006;20:501–13.


54. Roelfsema JH, Peters DJ. Rubinstein-Taybi syndrome: clinical and molecular overview. Expert Rev Mol Med 2007;9:1–16.

56. Maass PG, Aydin A, Luft FC, Schächterle C, Weise A, Stricker S, et al. PDE3A mutations cause autosomal dominant hypertension with brachydactyly. Nat Genet 2015;47:647–53.



57. Boda H, Uchida H, Takaiso N, Ouchi Y, Fujita N, Kuno A, et al. A PDE3A mutation in familial hypertension and brachydactyly syndrome. J Hum Genet 2016;61:701–3.



58. Vortkamp A, Lee K, Lanske B, Segre GV, Kronenberg HM, Tabin CJ. Regulation of rate of cartilage differentiation by Indian hedgehog and PTH-related protein. Science 1996;273:613–22.


59. Vasques GA, Funari MF, Ferreira FM, Aza-Carmona M, Sentchordi-Montané L, Barraza-García J, et al. IHH Gene Mutations causing short stature with nonspecific skeletal abnormalities and response to growth hormone therapy. J Clin Endocrinol Metab 2018;103:604–14.



60. Gao B, Guo J, She C, Shu A, Yang M, Tan Z, et al. Mutations in IHH, encoding Indian hedgehog, cause brachydactyly type A-1. Nat Genet 2001;28:386–8.



61. Hellemans J, Coucke PJ, Giedion A, De Paepe A, Kramer P, Beemer F, et al. Homozygous mutations in IHH cause acrocapitofemoral dysplasia, an autosomal recessive disorder with cone-shaped epiphyses in hands and hips. Am J Hum Genet 2003;72:1040–6.



62. Klopocki E, Hennig BP, Dathe K, Koll R, de Ravel T, Baten E, et al. Deletion and point mutations of PTHLH cause brachydactyly type E. Am J Hum Genet 2010;86:434–9.



63. Zhang P, Jobert AS, Couvineau A, Silve C. A homozygous inactivating mutation in the parathyroid hormone/parathyroid hormone-related peptide receptor causing Blomstrand chondrodysplasia. J Clin Endocrinol Metab 1998;83:3365–8.


64. Schipani E, Kruse K, Jüppner H. A constitutively active mutant PTH-PTHrP receptor in Jansen-type metaphyseal chondrodysplasia. Science 1995;268:98–100.


65. Potter LR, Abbey-Hosch S, Dickey DM. Natriuretic peptides, their receptors, and cyclic guanosine monophosphate-dependent signaling functions. Endocr Rev 2006;27:47–72.



66. Hisado-Oliva A, Ruzafa-Martin A, Sentchordi L, Funari MFA, Bezanilla-López C, Alonso-Bernáldez M, et al. Mutations in C-natriuretic peptide (NPPC): a novel cause of autosomal dominant short stature. Genet Med 2018;20:91–7.



67. Wang SR, Jacobsen CM, Carmichael H, Edmund AB, Robinson JW, Olney RC, et al. Heterozygous mutations in natriuretic peptide receptor-B (NPR2) gene as a cause of short stature. Hum Mutat 2015;36:474–81.



68. Moncla A, Missirian C, Cacciagli P, Balzamo E, Legeai-Mallet L, Jouve JL, et al. A cluster of translocation breakpoints in 2q37 is associated with overexpression of NPPC in patients with a similar overgrowth phenotype. Hum Mutat 2007;28:1183–8.


69. Miura K, Kim OH, Lee HR, Namba N, Michigami T, Yoo WJ, et al. Overgrowth syndrome associated with a gainof-function mutation of the natriuretic peptide receptor 2 (NPR2) gene. Am J Med Genet A 2014;164A:156–63.


70. Teixeira CC, Agoston H, Beier F. Nitric oxide, C-type natriuretic peptide and cGMP as regulators of endochondral ossification. Dev Biol 2008;319:171–8.



71. Krejci P, Masri B, Fontaine V, Mekikian PB, Weis M, Prats H, et al. Interaction of fibroblast growth factor and C-natriuretic peptide signaling in regulation of chondrocyte proliferation and extracellular matrix homeostasis. J Cell Sci 2005;118(Pt 21):5089–100.


72. Bartels CF, Bükülmez H, Padayatti P, Rhee DK, van Ravenswaaij-Arts C, Pauli RM, et al. Mutations in the transmembrane natriuretic peptide receptor NPR-B impair skeletal growth and cause acromesomelic dysplasia, type Maroteaux. Am J Hum Genet 2004;75:27–34.



73. Hachiya R, Ohashi Y, Kamei Y, Suganami T, Mochizuki H, Mitsui N, et sl. Intact kinase homology domain of natriuretic peptide receptor-B is essential for skeletal development. J Clin Endocrinol Metab 2007;92:4009–14.



74. Bocciardi R, Giorda R, Buttgereit J, Gimelli S, Divizia MT, Beri S, et al. Overexpression of the C-type natriuretic peptide (CNP) is associated with overgrowth and bone anomalies in an individual with balanced t(2;7) translocation. Hum Mutat 2007;28:724–31.


75. Miura K, Namba N, Fujiwara M, Ohata Y, Ishida H, Kitaoka T, et al. An overgrowth disorder associated with excessive production of cGMP due to a gain-of-function mutation of the natriuretic peptide receptor 2 gene. PLoS One 2012;7:e42180.



76. Hannema SE, van Duyvenvoorde HA, Premsler T, Yang RB, Mueller TD, Gassner B, et al. An activating mutation in the kinase homology domain of the natriuretic peptide receptor-2 causes extremely tall stature without skeletal deformities. J Clin Endocrinol Metab 2013;98:E1988–98.


77. Al-Qattan MM. Embryology of familial (non-syndromic) brachydactyly of the hand. J Hand Surg Eur Vol 2014;39:926–33.


78. Al-Qattan MM, Al-Motairi MI, Al Balwi MA. Two novel homozygous missense mutations in the GDF5 gene cause brachydactyly type C. Am J Med Genet A 2015;167:1621–6.


79. Farooq M, Nakai H, Fujimoto A, Fujikawa H, Kjaer KW, Baig SM, et al. Characterization of a novel missense mutation in the prodomain of GDF5, which underlies brachydactyly type C and mild Grebe type chondrodysplasia in a large Pakistani family. Hum Genet 2013;132:1253–64.



80. Martinez-Garcia M, Garcia-Canto E, Fenollar-Cortes M, Aytes AP, Trujillo-Tiebas MJ. Characterization of an acromesomelic dysplasia, Grebe type case: novel mutation affecting the recognition motif at the processing site of GDF5. J Bone Miner Metab 2016;34:599–603.



81. Thomas JT, Kilpatrick MW, Lin K, Erlacher L, Lembessis P, Costa T, et al. Disruption of human limb morphogenesis by a dominant negative mutation in CDMP1. Nat Genet 1997;17:58–64.



82. Chang SC, Hoang B, Thomas JT, Vukicevic S, Luyten FP, Ryba NJ, et al. Cartilage-derived morphogenetic proteins. New members of the transforming growth factor-beta superfamily predominantly expressed in long bones during human embryonic development. J Biol Chem 1994;269:28227–34.


83. Hatakeyama Y, Tuan RS, Shum L. Distinct functions of BMP4 and GDF5 in the regulation of chondrogenesis. J Cell Biochem 2004;91:1204–17.


84. Bai X, Xiao Z, Pan Y, Hu J, Pohl J, Wen J, et al. Cartilagederived morphogenetic protein-1 promotes the differentiation of mesenchymal stem cells into chondrocytes. Biochem Biophys Res Commun 2004;325:453–60.


85. Kuss P, Kraft K, Stumm J, Ibrahim D, Vallecillo-Garcia P, Mundlos S, et al. Regulation of cell polarity in the cartilage growth plate and perichondrium of metacarpal elements by HOXD13 and WNT5A. Dev Biol 2014;385:83–93.


86. Person AD, Beiraghi S, Sieben CM, Hermanson S, Neumann AN, Robu ME, et al. WNT5A mutations in patients with autosomal dominant Robinow syndrome. Dev Dyn 2010;239:327–37.



87. Roifman M, Marcelis CL, Paton T, Marshall C, Silver R, Lohr JL, et al. De novo WNT5A-associated autosomal dominant Robinow syndrome suggests specificity of genotype and phenotype. Clin Genet 2015;87:34–41.


88. Uchimura T, Hollander JM, Nakamura DS, Liu Z, Rosen CJ, Georgakoudi I, et al. An essential role for IGF2 in cartilage development and glucose metabolism during postnatal long bone growth. Development 2017;144:3533–46.



89. Begemann M, Zirn B, Santen G, Wirthgen E, Soellner L, Büttel HM, et al. Paternally Inherited IGF2 Mutation and Growth Restriction. N Engl J Med 2015;373:349–56.


90. Nilsson O, Marino R, De Luca F, Phillip M, Baron J. Endocrine regulation of the growth plate. Horm Res 2005;64:157–65.


91. Daughaday WH. Growth hormone axis overview--somatomedin hypothesis. Pediatr Nephrol 2000;14:537–40.



92. Fujimoto M, Kawashima Sonoyama Y, Hamajima N, Hamajima T, Kumura Y, Miyahara N, et al. Heterozygous nonsense mutations near the C-terminal region of IGF1R in two patients with small-for-gestational-age-related short stature. Clin Endocrinol (Oxf) 2015;83:834–41.


93. Abuzzahab MJ, Schneider A, Goddard A, Grigorescu F, Lautier C, Keller E, et al. IGF-I receptor mutations resulting in intrauterine and postnatal growth retardation. N Engl J Med 2003;349:2211–22.


94. Dauber A, Muñoz-Calvo MT, Barrios V, Domené HM, Kloverpris S, Serra-Juhé C, et al. Mutations in pregnancyassociated plasma protein A2 cause short stature due to low IGF-I availability. EMBO Mol Med 2016;8:363–74.



95. Bárcena C, Quesada V, De Sandre-Giovannoli A, Puente DA, Fernández-Toral J, Sigaudy S, et al. Exome sequencing identifies a novel mutation in PIK3R1 as the cause of SHORT syndrome. BMC Med Genet 2014;15:51.




96. Dyment DA, Smith AC, Alcantara D, Schwartzentruber JA, Basel-Vanagaite L, Curry CJ, et al. Mutations in PIK3R1 cause SHORT syndrome. Am J Hum Genet 2013;93:158–66.



97. Schroeder C, Riess A, Bonin M, Bauer P, Riess O, Döbler-Neumann M, et al. PIK3R1 mutations in SHORT syndrome. Clin Genet 2014;86:292–4.


98. Nilsson O, Weise M, Landman EB, Meyers JL, Barnes KM, Baron J. Evidence that estrogen hastens epiphyseal fusion and cessation of longitudinal bone growth by irreversibly depleting the number of resting zone progenitor cells in female rabbits. Endocrinology 2014;155:2892–9.




99. Verma N, Jain V, Birla S, Jain R, Sharma A. Growth and hormonal profile from birth to adolescence of a girl with aromatase deficiency. J Pediatr Endocrinol Metab 2012;25:1185–90.



100. Marino R, Perez Garrido N, Costanzo M, Guercio G, Juanes M, Rocco C, et al. Five new cases of 46,XX aromatase deficiency: clinical follow-up from birth to puberty, a novel mutation, and a founder effect. J Clin Endocrinol Metab 2015;100:E301–7.


101. Fukami M, Miyado M, Nagasaki K, Shozu M, Ogata T. Aromatase excess syndrome: a rare autosomal dominant disorder leading to pre- or peri-pubertal onset gynecomastia. Pediatr Endocrinol Rev 2014;11:298–305.

102. Quaynor SD, Stradtman EW Jr, Kim HG, Shen Y, Chorich LP, Schreihofer DA, et al. Delayed puberty and estrogen resistance in a woman with estrogen receptor α variant. N Engl J Med 2013;369:164–71.


103. Bernard V, Kherra S, Francou B, Fagart J, Viengchareun S, Guéchot J, et al. Familial multiplicity of estrogen insensitivity associated with a loss-of-function ESR1 mutation. J Clin Endocrinol Metab 2017;102:93–9.


104. Smith EP, Boyd J, Frank GR, Takahashi H, Cohen RM, Specker B, et al. Estrogen resistance caused by a mutation in the estrogen-receptor gene in a man. N Engl J Med 1994;331:1056–61.


105. Eggermann T. Epigenetic regulation of growth: lessons from Silver-Russell syndrome. Endocr Dev 2009;14:10–9.


106. Tatton-Brown K, Seal S, Ruark E, Harmer J, Ramsay E, Del Vecchio Duarte S, et al. Mutations in the DNA methyltransferase gene DNMT3A cause an overgrowth syndrome with intellectual disability. Nat Genet 2014;46:385–8.




107. Xin B, Cruz Marino T, Szekely J, Leblanc J, Cechner K, Sency V, et al. Novel DNMT3A germline mutations are associated with inherited Tatton-Brown-Rahman syndrome. Clin Genet 2017;91:623–8.


108. Tatton-Brown K, Loveday C, Yost S, Clarke M, Ramsay E, Zachariou A, et al. Mutations in epigenetic regulation genes are a major cause of overgrowth with intellectual disability. Am J Hum Genet 2017;100:725–36.



109. Gibson WT, Hood RL, Zhan SH, Bulman DE, Fejes AP, Moore R, et al. Mutations in EZH2 cause Weaver syndrome. Am J Hum Genet 2012;90:110–8.



110. Tatton-Brown K, Hanks S, Ruark E, Zachariou A, Duarte Sdel V, Ramsay E, et al. Germline mutations in the oncogene EZH2 cause Weaver syndrome and increased human height. Oncotarget 2011;2:1127–33.



111. Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P, et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 2002;298:1039–43.


112. Kurotaki N, Imaizumi K, Harada N, Masuno M, Kondoh T, Nagai T, et al. Haploinsufficiency of NSD1 causes Sotos syndrome. Nat Genet 2002;30:365–6.



113. Luscan A, Laurendeau I, Malan V, Francannet C, Odent S, Giuliano F, et al. Mutations in SETD2 cause a novel overgrowth condition. J Med Genet 2014;51:512–7.









