 |
 |
- Search
| Ann Pediatr Endocrinol Metab > Volume 22(2); 2017 > Article |
|
Abstract
Next-generation sequencing (NGS) and array-based comparative genomic hybridization (array CGH) have enabled us to perform high-throughput mutation screening and genome-wide copy number analysis, respectively. These methods can be used for molecular diagnosis of pediatric endocrine disorders. NGS has determined the frequency and phenotypic variation of mutations in several disease-associated genes. Furthermore, whole exome analysis using NGS has successfully identified several novel causative genes for endocrine disorders. Array CGH is currently used as the standard procedure for molecular cytogenetic analysis. Array CGH can detect various submicroscopic genomic rearrangements involving exons or enhancers of disease-associated genes. This review introduces some examples of the use of NGS and array CGH for the molecular diagnosis of pediatric endocrine disorders.
Single gene mutations and submicroscopic chromosomal rearrangements account for a certain percentage of the etiology of pediatric endocrine disorders1,2,3). Identification of such molecular defects is beneficial for patients, because molecular diagnosis often allows better clinical management and accurate genetic counseling4,5). Furthermore, identification of the genetic causes of congenital endocrine disorders provides advances in medical research.
Recently, various molecular technologies have been developed to identify genetic and genomic alterations in humans. These technologies include next generation sequencing (NGS) that can detect nucleotide substitutions and small insertions/deletions (indels)6,7,8), and array-based comparative genomic hybridization (array CGH) that identifies copy-number variations (CNVs) in the genome9,10,11,12). NGS and array CGH enable us to perform systematic molecular analyses for multiple clinical samples. This review introduces some examples of the use of NGS and array CGH for the molecular diagnosis of pediatric endocrine disorders.
NGS is a powerful tool for high-throughput mutation analysis6,7,8). This method is particularly useful to elucidate the genetic causes of Mendelian disorders with multiple causative genes. In our recent studies13,14,15,16), we performed NGS analyses for several samples of patients with pediatric endocrine disorders (Fig. 1). For example, we investigated the genetic causes of hypogonadotropic hypogonadism (HH), a rare condition caused by monoallelic, biallelic, or oligogenic mutations in more than 20 genes13,16). We created a custom-made NGS panel that covers most known causative genes for HH. DNA samples obtained from 58 patients with HH with and without additional clinical features were analyzed13). As a result, possible pathogenic mutations were identified in 13 of the 58 patients. Mutations in FGFR1 and CHD7 accounted for the majority of the molecular defects in our patients13). We identified an SOX3 polyalanine deletion, a rare mutation known to cause pituitary dysfunction17), in a patient with normosmic isolated HH (IHH)13). This finding expanded the phenotypic spectrum of SOX3 polyalanine deletion to include HH without other pituitary hormone deficiency. Likewise, we detected a WDR11 splice-site mutation in a patient with combined pituitary hormone deficiency (CPHD)13). These data indicated that haploinsufficiency of WDR11, a gene that has recently been implicated in normosmic IHH and Kallmann syndrome (HH with anosmia)18), can also underlie CPHD.
In another study, we analyzed 62 patients with nonsyndromic hypospadias (hypospadias without additional clinical features), a common anomaly in boys15). We created an NGS panel covering the coding exons of 25 known disease-associated genes, and detected mutations in AR, NR5A1, BNC2 and other genes in seven cases15). These data implied that more than 10% of cases with nonsyndromic hypospadias arise from mutations in known causative genes. Importantly, one patient carried probably damaging mutations in multiple genes15), suggesting the possible oligogenicity of nonsyndromic hypospadias.
The abovementioned results provided evidence that mutation screening using NGS is helpful to clarify the contribution of each causative gene to the etiology of polygenic disorders. In addition, NGS can provide critical information about phenotypic variations of each monogenic abnormality
NGS can also be used for systematic mutation screening of all known protein-coding genes (whole exome analysis)8,19). To date, whole exome analysis has successfully detected several novel causative genes for human disorders. For example, in 2013, Abreu et al.20) performed whole exome analysis of 15 patients with central precocious puberty (CPP) of unknown etiology. NGS analysis of the patients and their unaffected family members identified MKRN3 mutations cosegregated with the disease. Subsequent studies confirmed that MKRN3 mutations constitute one of the major genetic causes of CPP21,22,23).
Similarly, whole exome analysis helped to identify a novel cause of disorders of sex development (DSD). Bashamboo et al., Baetens et al., and our group revealed that an identical missense substitution in NR5A1, p.R92W, induces testicular development in genetic females24,25,26). This finding provided the first indication that specific missense mutations in a single gene can underlie 46,XX testicular/ovotesticular DSD. Since the p.R92W mutation in Nr5a1 did not induce testicular formation in female mice27), it appears that NR5A1 plays a species-specific role in sexual development.
Array CGH can screen CNVs in the entire genome in a single assay9,10,11,12). This method is currently used as the standard procedure for molecular cytogenetic analysis. Indeed, array CGH has higher resolution than conventional G-banding12), although array CGH cannot detect copy-number neutral translocations and inversions. To date, it is known that CNVs account for a significant fraction of the etiology of Mendelian disorders12,28). We have performed array CGH for several patients with pediatric endocrine disorders14,29). We analyzed DNA samples obtained from 24 patients with 46,XY DSD of unknown etiology, and identified submicroscopic deletions in 3 patients29). These results suggested that submicroscopic CNVs represent important genetic causes of 46,XY DSD. These deletions were predicted to induce undermasculinized external genitalia by eliminating exons or cis-regulatory elements of genes involved in sex development. Notably, one patient with complete 46,XY gonadal dysgenesis was found to carry a microdeletion in a far upstream region of SOX9 (Fig. 2)14), indicating that a testis-specific enhancer(s) of SOX9 is located within this deleted region.
Similarly, we utilized array CGH to characterize CNVs in the short arm pseudoautosomal region of sex chromosomes (PAR1) associated with idiopathic short stature or Leri-Weill dyschondrosteosis (short stature accompanied by Madelung deformity in the forearm)30,31,32). It is known that CNVs in PAR1 encompassing SHOX and/or its putative enhancers are important causes of short stature31,33,34). These CNVs were predicted to affect skeletal growth by reducing SHOX expression levels in chondrocytes30,31). We found that CNVs in PAR1 are highly variable in their sizes and positions30,31,32). Breakpoint characterization of the CNVs suggested that various mechanisms such as non-allelic homologous recombination and non-homologous end joining, and possibly replication errors as well, are involved in these abnormalities30,31,34,35). Thus, SHOX abnormality represents a model of human genomic disorders.
Array CGH has also identified novel genomic abnormalities associated with pediatric endocrine disorders. For example, we performed array CGH analysis for patients with hereditary gynecomastia (aromatase excess syndrome), a rare autosomal dominant disorder characterized by estrogen overproduction due to excessive enzymatic activity of aromatase36,37,38,39). We found that these patients have submicroscopic CNVs in the upstream region of the aromatase gene (CYP19A1)36,37,38,39). These CNVs included microduplications encompassing CYP19A1 promoters and microdeletions that lead to aberrant fusion between the coding exons of CYP19A1 and promoters of widely expressed neighboring genes38). These data suggested that hereditary gynecomastia is caused by submicroscopic genomic rearrangements leading to CYP19A1 overexpression38). Most importantly, our results indicated for the first time that a tandem duplication involving physiological promoters can underlie gene overexpression and resultant congenital disorders36,38).
Although the use of NGS and array CGH is highly beneficial for patients with pediatric endocrine disorders as well as other genetic diseases, these new technologies also raise ethical issues that are not associated with conventional molecular analyses. For example, whole exome analysis can detect mutations that are not related with the patient's present phenotype40,41). Some of these unexpected genetic alterations may have a potential to cause some serious disorders later in life. In addition, NGS and array CGH frequently detect molecular alterations of unknown pathogenicity 5). There is debate as to which and how the findings of systematic molecular analyses should be reported to the patients. The American College of Medical Genetics and Genomics and other organizations have published recommendations for the management of data obtained from NGS analysis41,42,43).
In addition, NGS and array CGH can be applied to prenatal diagnosis44,45). For example, NGS is used for noninvasive prenatal testing for chromosomal aneuploidy45). NGS and array CGH have broad applicability in the prenatal molecular diagnosis of both Mendelian disorders and multifactorial conditions5). However, the benefits, limitations, and ethical issues of these technologies need to be discussed.
NGS and array CGH are beneficial both for clinical practice and for research on pediatric endocrinology. In particular, since the cost of NGS is continuously decreasing, the utility of this method will be further enhanced in the future. Moreover, NGS has additional applications such as characterization of gut microbial flora in individuals with specific conditions46,47). The use of NGS and array CGH will provide further information about the causative mechanisms of pediatric endocrine disorders.
Acknowledgments
This study was supported by the Grants-in-Aid from the Japan Society for the Promotion of Science; and by the Grants from the Ministry of Health, Labor and Welfare, the Japan Agency for Medical Research and Development, the National Center for Child Health and Development and the Takeda Foundation.
References
1. Lee C, Iafrate AJ, Brothman AR. Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat Genet 2007;39(7 Suppl):S48ŌĆōS54. PMID: 17597782.


2. Buonocore F, Achermann JC. Human sex development: targeted technologies to improve diagnosis. Genome Biol 2016;17:257. PMID: 27978845.




3. Eggers S, Sadedin S, van den Bergen JA, Robevska G, Ohnesorg T, Hewitt J, et al. Disorders of sex development: insights from targeted gene sequencing of a large international patient cohort. Genome Biol 2016;17:243. PMID: 27899157.




5. Katsanis SH, Katsanis N. Molecular genetic testing and the future of clinical genomics. Nat Rev Genet 2013;14:415ŌĆō426. PMID: 23681062.



6. Smith DR, Quinlan AR, Peckham HE, Makowsky K, Tao W, Woolf B, et al. Rapid whole-genome mutational profiling using next-generation sequencing technologies. Genome Res 2008;18:1638ŌĆō1642. PMID: 18775913.



7. Shendure J, Ji H. Next-generation DNA sequencing. Nat Biotechnol 2008;26:1135ŌĆō1145. PMID: 18846087.


8. Metzker ML. Sequencing technologies: the next generation. Nat Rev Genet 2010;11:31ŌĆō46. PMID: 19997069.


9. Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D, et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998;20:207ŌĆō211. PMID: 9771718.


10. Vissers LE, de Vries BB, Osoegawa K, Janssen IM, Feuth T, Choy CO, et al. Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 2003;73:1261ŌĆō1270. PMID: 14628292.



11. Feuk L, Carson AR, Scherer SW. Structural variation in the human genome. Nat Rev Genet 2006;7:85ŌĆō97. PMID: 16418744.


12. Miller DT, Adam MP, Aradhya S, Biesecker LG, Brothman AR, Carter NP, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet 2010;86:749ŌĆō764. PMID: 20466091.



13. Izumi Y, Suzuki E, Kanzaki S, Yatsuga S, Kinjo S, Igarashi M, et al. Genome-wide copy number analysis and systematic mutation screening in 58 patients with hypogonadotropic hypogonadism. Fertil Steril 2014;102:1130ŌĆō1136.e3. PMID: 25064402.


14. Katoh-Fukui Y, Igarashi M, Nagasaki K, Horikawa R, Nagai T, Tsuchiya T, et al. Testicular dysgenesis/regression without campomelic dysplasia in patients carrying missense mutations and upstream deletion of SOX9. Mol Genet Genomic Med 2015;3:550ŌĆō557. PMID: 26740947.



15. Kon M, Suzuki E, Dung VC, Hasegawa Y, Mitsui T, Muroya K, et al. Molecular basis of non-syndromic hypospadias: systematic mutation screening and genome-wide copy-number analysis of 62 patients. Hum Reprod 2015;30:499ŌĆō506. PMID: 25605705.



16. Fukami M, Suzuki E, Izumi Y, Torii T, Narumi S, Igarashi M, et al. Paradoxical gain-of-function mutant of the G-protein-coupled receptor PROKR2 promotes early puberty. J Cell Mol Med 2017;3 24 [Epub]. PMID: 10.1111/jcmm.13146.



17. Alatzoglou KS, Kelberman D, Cowell CT, Palmer R, Arnhold IJ, Melo ME, et al. Increased transactivation associated with SOX3 polyalanine tract deletion in a patient with hypopituitarism. J Clin Endocrinol Metab 2011;96:E685ŌĆōE690. PMID: 21289259.



18. Kim HG, Ahn JW, Kurth I, Ullmann R, Kim HT, Kulharya A, et al. WDR11, a WD protein that interacts with transcription factor EMX1, is mutated in idiopathic hypogonadotropic hypogonadism and Kallmann syndrome. Am J Hum Genet 2010;87:465ŌĆō479. PMID: 20887964.



19. Bamshad MJ, Ng SB, Bigham AW, Tabor HK, Emond MJ, Nickerson DA, et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet 2011;12:745ŌĆō755. PMID: 21946919.


20. Abreu AP, Dauber A, Macedo DB, Noel SD, Brito VN, Gill JC, et al. Central precocious puberty caused by mutations in the imprinted gene MKRN3. N Engl J Med 2013;368:2467ŌĆō2475. PMID: 23738509.



21. Settas N, Dacou-Voutetakis C, Karantza M, Kanaka-Gantenbein C, Chrousos GP, Voutetakis A. Central precocious puberty in a girl and early puberty in her brother caused by a novel mutation in the MKRN3 gene. J Clin Endocrinol Metab 2014;99:E647ŌĆōE651. PMID: 24438377.



22. Shin YL. An update on the genetic causes of central precocious puberty. Ann Pediatr Endocrinol Metab 2016;21:66ŌĆō69. PMID: 27462581.



23. Christoforidis A, Skordis N, Fanis P, Dimitriadou M, Sevastidou M, Phelan MM, et al. A novel MKRN3 nonsense mutation causing familial central precocious puberty. Endocrine 2017;56:446ŌĆō449. PMID: 28132164.



24. Bashamboo A, Donohoue PA, Vilain E, Rojo S, Calvel P, Seneviratne SN, et al. A recurrent p.Arg92Trp variant in steroidogenic factor-1 (NR5A1) can act as a molecular switch in human sex development. Hum Mol Genet 2016;25:3446ŌĆō3453. PMID: 27378692.



25. Baetens D, Stoop H, Peelman F, Todeschini AL, Rosseel T, Coppieters F, et al. NR5A1 is a novel disease gene for 46,XX testicular and ovotesticular disorders of sex development. Genet Med 2017;19:367ŌĆō376. PMID: 27490115.



26. Igarashi M, Takasawa K, Hakoda A, Kanno J, Takada S, Miyado M, et al. Identical NR5A1 missense mutations in two unrelated 46,XX individuals with testicular tissues. Hum Mutat 2017;38:39ŌĆō42. PMID: 27610946.


27. Miyado M, Inui M, Igarashi M, Katoh-Fukui Y, Takasawa K, Hakoda A, et al. The p.R92W variant of NR5A1/Nr5a1 induces testicular development of 46,XX gonads in humans, but not in mice: phenotypic comparison of human patients and mutation-induced mice. Biol Sex Differ 2016;7:56. PMID: 27833742.



28. Beckmann JS, Estivill X, Antonarakis SE. Copy number variants and genetic traits: closer to the resolution of phenotypic to genotypic variability. Nat Rev Genet 2007;8:639ŌĆō646. PMID: 17637735.


29. Igarashi M, Dung VC, Suzuki E, Ida S, Nakacho M, Nakabayashi K, et al. Cryptic genomic rearrangements in three patients with 46,XY disorders of sex development. PLoS One 2013;8:e68194. PMID: 23861871.



30. Fukami M, Naiki Y, Muroya K, Hamajima T, Soneda S, Horikawa R, et al. Rare pseudoautosomal copy-number variations involving SHOX and/or its flanking regions in individuals with and without short stature. J Hum Genet 2015;60:553ŌĆō556. PMID: 26040210.


31. Fukami M, Seki A, Ogata T. SHOX haploinsufficiency as a cause of syndromic and nonsyndromic short stature. Mol Syndromol 2016;7:3ŌĆō11. PMID: 27194967.



32. Shima H, Tanaka T, Kamimaki T, Dateki S, Muroya K, Horikawa R, et al. Systematic molecular analyses of SHOX in Japanese patients with idiopathic short stature and Leri-Weill dyschondrosteosis. J Hum Genet 2016;61:585ŌĆō591. PMID: 26984564.


33. Benito-Sanz S, Barroso E, Heine-Su├▒er D, Hisado-Oliva A, Romanelli V, Rosell J, et al. Clinical and molecular evaluation of SHOX/PAR1 duplications in Leri-Weill dyschondrosteosis (LWD) and idiopathic short stature (ISS). J Clin Endocrinol Metab 2011;96:E404ŌĆōE412. PMID: 21147883.



34. Benito-Sanz S, Royo JL, Barroso E, Paumard-Hern├Īndez B, Barreda-Bonis AC, Liu P, et al. Identification of the first recurrent PAR1 deletion in L├®ri-Weill dyschondrosteosis and idiopathic short stature reveals the presence of a novel SHOX enhancer. J Med Genet 2012;49:442ŌĆō450. PMID: 22791839.


35. Fukami M, Kato F, Tajima T, Yokoya S, Ogata T. Transactivation function of an approximately 800-bp evolutionarily conserved sequence at the SHOX 3' region: implication for the downstream enhancer. Am J Hum Genet 2006;78:167ŌĆō170. PMID: 16385461.



36. Fukami M, Shozu M, Soneda S, Kato F, Inagaki A, Takagi H, et al. Aromatase excess syndrome: identification of cryptic duplications and deletions leading to gain of function of CYP19A1 and assessment of phenotypic determinants. J Clin Endocrinol Metab 2011;96:E1035ŌĆōE1043. PMID: 21470988.


37. Fukami M, Tsuchiya T, Vollbach H, Brown KA, Abe S, Ohtsu S, et al. Genomic basis of aromatase excess syndrome: recombination- and replication-mediated rearrangements leading to CYP19A1 overexpression. J Clin Endocrinol Metab 2013;98:E2013ŌĆōE2021. PMID: 24064691.



38. Fukami M, Miyado M, Nagasaki K, Shozu M, Ogata T. Aromatase excess syndrome: a rare autosomal dominant disorder leading to pre- or peri-pubertal onset gynecomastia. Pediatr Endocrinol Rev 2014;11:298ŌĆō305. PMID: 24716396.

39. Shihara D, Miyado M, Nakabayashi K, Shozu M, Ogata T, Nagasaki K, et al. Aromatase excess syndrome in a family with upstream deletion of CYP19A1. Clin Endocrinol (Oxf) 2014;81:314ŌĆō316. PMID: 24102311.


40. Vasli N, B├Čhm J, Le Gras S, Muller J, Pizot C, Jost B, et al. Next generation sequencing for molecular diagnosis of neuromuscular diseases. Acta Neuropathol 2012;124:273ŌĆō283. PMID: 22526018.



41. Green RC, Berg JS, Grody WW, Kalia SS, Korf BR, Martin CL, et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med 2013;15:565ŌĆō574. PMID: 23788249.



42. Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405ŌĆō424. PMID: 25741868.



43. Matthijs G, Souche E, Alders M, Corveleyn A, Eck S, Feenstra I, et al. Guidelines for diagnostic next-generation sequencing. Eur J Hum Genet 2016;24:2ŌĆō5. PMID: 26508566.



44. Wapner RJ, Martin CL, Levy B, Ballif BC, Eng CM, Zachary JM, et al. Chromosomal microarray versus karyotyping for prenatal diagnosis. N Engl J Med 2012;367:2175ŌĆō2184. PMID: 23215555.



45. Benn P, Cuckle H, Pergament E. Non-invasive prenatal testing for aneuploidy: current status and future prospects. Ultrasound Obstet Gynecol 2013;42:15ŌĆō33. PMID: 23765643.


46. Marrs T, Flohr C. The role of skin and gut microbiota in the development of atopic eczema. Br J Dermatol 2016;175(Suppl 2):13ŌĆō18.

47. Ma L, Xie Y, Han Z, Giesy JP, Zhang X. Responses of earthworms and microbial communities in their guts to Triclosan. Chemosphere 2017;168:1194ŌĆō1202. PMID: 27810239.


Fig.┬Ā1
Representative results of next generation sequencing. Each gray bar denotes a 100-bp DNA fragment obtained from a patient's sample. Substituted nucleotides are highlighted by colored letters. This patient carries a heterozygous A to C substitution (the blue box). All other substitutions in this figure are likely to be sequence errors.
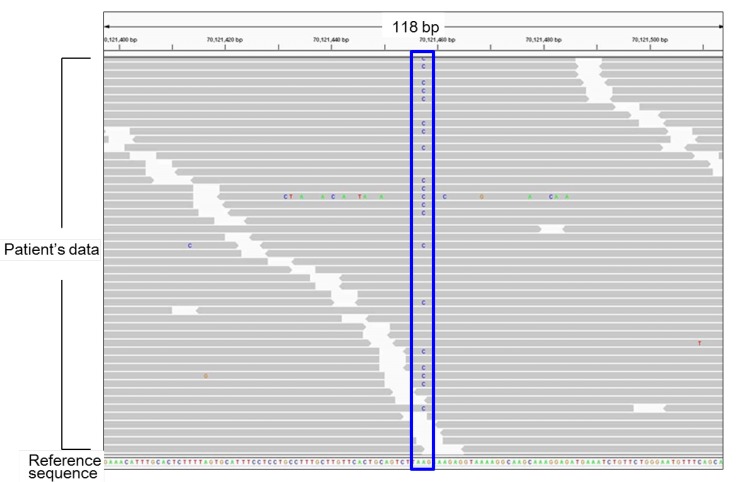
Fig.┬Ā2
Representative results of array-based comparative genomic hybridization. Submicroscopic deletion identified in a patient with 46,XY disorders of sex development is shown. This deletion likely encompasses a cis-acting enhancer(s) of SOX914). The deletion was shared by the patient's mother. The black, red, and green dots denote signals indicative of the normal, the increased (>+0.5) and the decreased (<ŌłÆ1.0) copy-numbers, respectively. Green arrows depict deleted regions. Genomic positions refer to the Human Genome (hg19, build 37).








