 |
 |
- Search
| Ann Pediatr Endocrinol Metab > Volume 21(3); 2016 > Article |
|
Abstract
Exosomes are extracellular vesicles that contain molecules that regulate the metabolic functions of adjacent or remote cells. Recent in vitro, in vivo and clinical studies support the hypothesis that exosomes released from various cell types play roles in the progression of metabolic disorders including type 2 diabetes. Based on this concept and advances in other diseases, the proteins, mRNA, microRNA and lipids in exosomes isolated from biological fluids have been proposed as biomarkers in metabolic disorders. However, several problems with the development of clinically applicable biomarkers have not been resolved. In this review, the biologic functions of exosomes are briefly introduced, and we discuss the technical and practical pros and cons of different methods of exosome isolation for the identification of exosomal biomarkers of metabolic disorders. Standardization of preanalytical variables and isolation of high-purity exosomes from fully characterized biological fluids will be necessary for the identification of useful exosomal biomarkers that can provide insights into the pathogenic mechanisms of complications of metabolic syndrome and of whole-body metabolism.
Exosomes are a class of extracellular vesicles (EVs), sized around 100 nm (30–120 nm) in diameter, that are released from many different cell types. Exosomes differ from another class of EVs, microvesicles (MVs, also referred to as 'ectosomes') that are of a different size (up to 2 µm) and are produced by a different mechanism. Exosomes were first discovered over 30 years ago and were considered to be little more than a means of cellular garbage disposal. However, more recent studies have provided scientific evidence that exosomes play roles in intercellular communication. Advances in the study of exosomes have identified the mechanisms regulating their production and have shown that exosomes contain cell-type specific collections of proteins, lipids and genetic material that can be transferred to recipient cells where they regulate function. Recently, scientific attention has focused on the various functions of exosomes in the pathophysiology of diseases including cancer, metabolic diseases, cardiovascular diseases, immune diseases, and neurodegenerative diseases. In particular, it has been shown that in type 2 diabetes mellitus (T2DM), exosomes secreted from skeletal muscle, visceral adipose tissue (VAT) and hepatocytes can transfer both functional proteins and RNA species that regulate the metabolic function of both remote tissues and of adjacent cells1,2,3).
The range of biomolecules in exosomes is diverse, including soluble and membrane-bound proteins, lipids, mRNA, microRNAs (miRNAs) and noncoding RNAs (ncRNAs). Therefore, exosomes are potentially a good source of disease biomarkers for early diagnosis and/or prediction of disease progression. However, there are several unresolved problems with the isolation of exosomes from biological fluids. In this review, we describe the potential roles of exosomes from diverse cell types in the regulation of cell metabolism, and discuss the possibility of the clinical use of blood-based exosomal biomarkers and the challenges for the use of exosomes as a source of disease biomarkers.
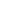
Exosomes originate from sections of the plasma membrane that are heavily enriched in phosphatidylserine. As illustrated in Fig. 1, the first step of exosome formation is inward budding of the plasma membrane to form an early endosome. During maturation of the endosome, the inward budding of limited areas of the endosomal membrane to form nano-sized vesicles produces multivesicular bodies (MVBs) that contain intraluminal vesicles (ILVs), in which cytoplasmic components including soluble proteins and nucleic acids are present. Although the mechanisms by which proteins are loaded into exosomes are not fully understood, the endosomal sorting complexes required for transport machinery is essential for the formation of ILVs in an MVB4,5). The inward budding of ILVs and the sorting of cytosolic proteins or nucleic acids also involve tetraspanin- and ceramide-dependent interactions6). Once MVBs are formed, the ILVs (exosomes) are released to the extracellular environment by fusion of the MVBs with the plasma membrane, a process that is regulated by a small GTPase-dependent mechanism. Alternatively, MVBs can fuse with a lysosome or autophagosome, where the contents of the MVBs are degraded (Fig. 1). The secreted exosomes function in fundamental pleiotropic biologic processes via direct interaction of their transmembrane proteins or lipid ligands with cell surface receptors, or by delivering their contents (e.g., transcription factors, cytosolic proteins, mRNA, miRNA and ncRNA) into recipient cells via membrane fusion7). Their biologic effects can be local (such as regulation of synaptic plasticity) or effected by delivery of secreted exosomes to remote target cells. These physiologic effects include, but are not limited to, the regulation of immune and neuronal function8,9), cell phenotype modulation10,11) and tissue regeneration12,13). It is not surprising that exosomes also have significant pathological roles in a variety of diseases including tumor progression14,15), promoting immune escape16), transmission of neuropathological proteins17,18), and metabolic dysfunction19).
Although studies that clearly demonstrate the nature of EVs are limited, the number of EVs is increased in the plasma of patients with insulin resistance and T2DM20,21). In particular, lipids or surface molecules of EVs released from skeletal muscle, platelets, or T cells increase the risk of metabolic dysfunction or atherogenic tendency19,20,21). In addition, an increased number of EVs has also been linked with obesity, micro/macrovascular complications, and inflammatory conditions22,23,24,25), indicating that the quantity of exosomes can be an indicator of these diseases. Recent studies demonstrated that exosomes can orchestrate insulin signaling in peripheral tissues, which suggested the emerging concept of exosome-mediated regulation of metabolism in T2DM19,26,27,28,29). For example, exosomes released from VAT in diabetic mice can fuse with blood monocytes and stimulate their differentiation into activated macrophages30). Exosomes released from VAT in mice with T2DM also induced the development of insulin resistance when they were administered to normal mice30). Adipocyte-derived exosomes contain hundreds of proteins including adipokines28), which may indicate a role for exosomes in the intercellular transmission of signals. Exosomes released from skeletal muscle, another critical tissue regulating whole-body metabolism, can be incorporated into various tissues including skeletal muscle, pancreas and liver, and can modify muscle homeostasis19).
Based on these findings and advances in other diseases, attempts are being made to develop novel diagnostic or prognostic biomarkers using exosomal molecules isolated from biological fluids. For example, the presence of diseaseor stage-specific miRNAs in blood exosomes may be a reliable and clinically applicable candidate biomarker for diagnosis or prediction of various diabetic complications31,32,33).
A biomarker is defined as 'a characteristic that is objectively measured and evaluated as an indicator of a normal biologic process, pathogenic process, or pharmacologic response to a therapeutic intervention'34). Exosomes can be detected in a variety of biological fluids including serum, plasma, urine, saliva, cerebrospinal fluid (CSF) and milk. In particular, serumor plasma-derived exosomes are a promising avenue to be explored for biomarker development. T2DM is the principal component of metabolic syndrome (MetS); therefore, based on the known pathophysiologic processes of MetS, blood-based biomarkers of T2DM could offer a minimally invasive method to predict patients' outcomes, a modality for predicting the response to therapeutic intervention, and an opportunity to provide earlier care to avoid the development of more severe complications. Patients with different types of diabetes and different complications of diabetes showed diverse patterns of exosomal biomolecules, suggesting that exosomes may contribute to tissue- or stage-specific pathogenic mechanisms of diabetes and its complications35). Exosomes from specific tissues including liver, muscle, endothelium, blood cells and VAT can travel in the bloodstream and be distributed to remote tissues. In the target tissues, exosomes that are derived from remote or adjacent tissues integrate with or interact with recipient cells, and act to regulate gene transcription, posttranslational modification, signal transduction, and metabolism. In fact, white VAT, an important organ that regulates whole-body metabolism and metabolic signaling and modulates the function of remote tissues including liver36) and macrophages30), is known to release exosomes. Notably, VAT dysfunction is an independent risk factor for cardiovascular complications of T2DM37,38). The pathophysiological mechanisms that link cardiovascular complications and VAT dysfunction are still poorly understood, but blood-based biomarkers that indicated the risk of diabetes complications would be extremely useful. Because diverse cellular cargoes are delivered by exosomes, these could be a source of blood-based biomarkers. Therefore, exosomes isolated from plasma or serum might present a molecular signature of a combination of tissue types, and could represent both common and specific markers of diverse cells and tissues or disease status. In contrast, exosomes from other types of biological fluids such as CSF, milk, or saliva may be tissue specific. The matrix of the biological fluid may also influence the exosome enrichment procedure; for example, albumin in plasma can be a major protein contaminant of isolated exosomes. Therefore, the procedure by which exosomes are isolated from biological fluids is critical for the development of exosomal biomarkers using various types of biological fluid.
The characterization of samples for exosome enrichment is the initial and essential step for exosomal biomarker development, because the extracellular milieu is complex, and plasma and serum, in particular, are biological fluids that reflect the extracellular environment of entire organs and of endothelium and blood cells. Therefore, the efficiency and recovery of exosomes isolated from biological fluids and the contaminants in the isolated exosomes are dependent on the matrix of the sample. There are several proposed methods to enrich exosomes from biological fluids and in vitro cell culture medium, but the optimal method of exosome isolation has not been established. The most widely used method for exosome enrichment is differential centrifugation, in which sequential rounds of centrifugation are intended to pellet cellular debris (10 minutes at 300 g), large vesicles (10 minutes at 2,000 g), MVs larger than 150 nm (30 minutes at 10,000 g) and exosomes (ultracentrifugation for 70 minutes at 100,000 g). Washing the final pellet that is enriched with exosomes can reduce its contamination by soluble proteins.
However, there are several issues with standardized exosome isolation using the differential centrifugation method. First, the duration of each centrifugation must be optimized according to the rotor type used. Two types of rotors are commonly used: fixed-angle or swing-bucket rotors. Because the sedimentation efficiency of EVs of diverse size and density is dependent on the sedimentation path length, the duration of centrifugation must be calculated for each rotor type39). Second, contamination of exosomes prepared by ultracentrifugation with small nonexosomal vesicles must be considered. Indeed, MVs that are generated by blebbing from plasma membranes have a broad size range (50–2,000 nm) that overlaps with the size of exosomes40,41). Therefore, we cannot exclude the possibility that a disease-specific exosomal biomarker is actually a combination of biomolecules from exosomes and small MVs. To exclude this possibility, in addition to size-based characterization the isolated exosomes should be characterized using at least three exosomal markers (e.g., tetraspanins, TSG101, and Alix) and the absence of or low levels of contamination with nonexosomal markers (e.g., Grp94, GM130, and cytochrome C) should be confirmed42). Exosomes can be further characterized using a combination of immunodetection of exosomal markers with electron microscopy (EM). EM is valuable for assessments of the morphology and size of vesicles in the samples (Fig. 2). Third, ultracentrifugation can induce the collapse and damage of exosomal membranes or aggregation of exosomes. Damage to the exosomal membrane can influence the quantification of biomarker levels. The formation of aggregates composed of a mixture of EVs of various phenotypes and morphologies may lead to misinterpretation of the level of exosomal biomarkers43). Finally, the conditions for differential centrifugation should be optimized according to the specific characteristics of each biological fluid (e.g., the difference in viscosity between plasma and urine).
Exosomes can also be purified by density gradient (DG) isolation with iodixanol (OptiPrep, Sigam-Aldrich, St. Louis, MO, USA) using a sucrose gradient followed by ultracentrifugation. Ultracentrifugation of samples overlaid on an iodixanol gradient for 16 hours at 100,000 g produces a layer that contains exosomes, which corresponds to a buoyant density of 1.13–1.19 g/mL. This method can prepare a relatively homogeneous exosome fraction with low levels of contamination44,45), and may be the best method for preparing highly purified exosomes for in vitro experiments. However, the preparation of exosomes by DG is relatively labor intensive, poorly reproducible and hard to standardize for routine clinical application.
Another method to prepare exosomes is size-exclusion chromatography (SEC). One or several fractions of a sample are loaded onto a packed SEC column that allows passage of nondamaged vesicles of regular shape and homogeneous size46). Larger vesicles can enter fewer pores in the column than can the smaller vesicles. The main advantages of this procedure for clinical applicability are its simplicity, good reproducibility and minimal damage to vesicles. However, this method cannot exclude the possibility of protein or RNA contamination, and because the sample volume should not exceed 10% of the resin volume the SEC method is less useful for larger-volume samples (e.g., urine or cell culture media) because of the cost. This limitation can be partially resolved by combining SEC with the use of commercially available ultrafiltration units (e.g., Vivaspin [Sartorius, Bohemia, NY, USA] or Amicon [Millipore, Darmstadt, Germany]) that keep exosomes on the filter surface and allow the removal of the majority of the sample fluid.
Finally, commercially available kits for polymer-based precipitation or immune bead-based capture methods (e.g., ExoQuick [System Biosciences, Palo Alto, CA, USA], Exo-spin [Cell Guidance Systems, St. Louis, MO, USA]) have been used in several biomarker studies to isolate exosomes from plasma47,48). The polymer-based precipitation kits enable high-throughput, fast and simple isolation of exosomes from low-volume samples. However, it should be noted that the purity of the exosomes was inferior to that obtained by other methods49) and that contamination by extracellular proteins was a major concern. The immune bead-based capture method produces a highly selected subpopulation of vesicles that is specific to a certain population or disease status and may not be generalizable. In addition, the availability of suitable antibodies for exosome isolation may be a limiting factor.
Compared with healthy controls, there were significantly more plasma EVs in patients with MetS50,51), supporting the hypothesis that quantification of EVs in blood may have potential as a screening biomarker for MetS. There are limited data on the clinical applicability of exosomal biomarkers for prediction of T2DM progression. However, considering the biologic roles of the molecular contents of exosomes in regulation of tissue-specific and/or whole-body metabolism, several studies have supported the hypothesis that proteins or miRNA in circulating or urinary exosomes (or microparticles) may be potential biomarkers to predict disease progression. For example, certain proteins and miRNAs in circulating or urinary EVs are associated with cardiometabolic risk factors21,23,29,52,53,54) and renal complications of diabetes55,56,57), respectively. In addition, these exosomal biomarkers may be useful for monitoring the pharmacological or therapeutic responses of patients with MetS and T2DM. For example, the levels of miRNAs let-7a and let-7f in circulating EVs were normalized by oral antidiabetes medication29), although it is unclear whether these changes were the result of pharmacological regulation of the underlying pathophysiology or were merely related to an improvement of glucose control.
These clinical studies provided molecular evidence that exosomes in biological fluids are good sources of potential biomarkers for metabolic diseases. There are currently no biomarkers of circulating or urinary exosomes that are validated for use in routine clinical practice. Nevertheless, although the analysis of circulating or urinary exosomes in metabolic disease is currently limited, exosomes may be a good source of diagnostic and prognostic biomarkers. To this end, longitudinal clinical studies with large sample size and with standardized system for the analysis of exosomal cargos should be conducted to evaluate clinical values of exosomal biomarkers for prediction of the progression of metabolic diseases.
In conclusion, there is increasing evidence that exosomes play roles in the regulation of tissue-specific and/or whole-body metabolism via the delivery of diverse exosomal cargo molecules to adjacent and remote tissues. Recent advances in the understanding of the biologic functions of exosomes facilitate clinical research towards the identification of biomarkers by using several methods to isolate exosomes from biological fluids. However, global efforts towards the standardization of preanalytical factors and procedures for exosome isolation (e.g., by the International Society for Extracellular Vesicles) are critical to facilitate the identification of clinically applicable exosomal biomarkers and overcome current challenges58). In addition, the characteristics of specific biological fluids used for exosome isolation must be considered (e.g., the effects of the different viscosities and volumes of plasma and urine samples). The development of a less labor intensive, cheap, high-throughput and valid method for purifying exosomes from biological fluids is a prerequisite for the identification of clinically applicable exosomal biomarkers.
References
1. Guescini M, Guidolin D, Vallorani L, Casadei L, Gioacchini AM, Tibollo P, et al. C2C12 myoblasts release micro-vesicles containing mtDNA and proteins involved in signal transduction. Exp Cell Res 2010;316:1977–1984. PMID: 20399774.


2. Aoki N, Jin-no S, Nakagawa Y, Asai N, Arakawa E, Tamura N, et al. Identification and characterization of microvesicles secreted by 3T3-L1 adipocytes: redox- and hormone-dependent induction of milk fat globule-epidermal growth factor 8-associated microvesicles. Endocrinology 2007;148:3850–3862. PMID: 17478559.



3. Conde-Vancells J, Rodriguez-Suarez E, Embade N, Gil D, Matthiesen R, Valle M, et al. Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J Proteome Res 2008;7:5157–5166. PMID: 19367702.



4. Raiborg C, Stenmark H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature 2009;458:445–452. PMID: 19325624.


5. Baietti MF, Zhang Z, Mortier E, Melchior A, Degeest G, Geeraerts A, et al. Syndecan-syntenin-ALIX regulates the biogenesis of exosomes. Nat Cell Biol 2012;14:677–685. PMID: 22660413.


6. Villarroya-Beltri C, Baixauli F, Gutiérrez-Vázquez C, Sánchez-Madrid F, Mittelbrunn M. Sorting it out: regulation of exosome loading. Semin Cancer Biol 2014;28:3–13. PMID: 24769058.



7. EL Andaloussi S, Mäger I, Breakefield XO, Wood MJ. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat Rev Drug Discov 2013;12:347–357. PMID: 23584393.


8. Simhadri VR, Reiners KS, Hansen HP, Topolar D, Simhadri VL, Nohroudi K, et al. Dendritic cells release HLA-B-associated transcript-3 positive exosomes to regulate natural killer function. PLoS One 2008;3:e3377. PMID: 18852879.



9. Chivet M, Hemming F, Pernet-Gallay K, Fraboulet S, Sadoul R. Emerging role of neuronal exosomes in the central nervous system. Front Physiol 2012;3:145. PMID: 22654762.



10. Jang YY, Collector MI, Baylin SB, Diehl AM, Sharkis SJ. Hematopoietic stem cells convert into liver cells within days without fusion. Nat Cell Biol 2004;6:532–539. PMID: 15133469.


11. Quesenberry PJ, Aliotta JM. Cellular phenotype switching and microvesicles. Adv Drug Deliv Rev 2010;62:1141–1148. PMID: 20558219.



12. Lai RC, Chen TS, Lim SK. Mesenchymal stem cell exosome: a novel stem cell-based therapy for cardiovascular disease. Regen Med 2011;6:481–492. PMID: 21749206.


13. Timmers L, Lim SK, Hoefer IE, Arslan F, Lai RC, van Oorschot AA, et al. Human mesenchymal stem cell-conditioned medium improves cardiac function following myocardial infarction. Stem Cell Res 2011;6:206–214. PMID: 21419744.


14. Camussi G, Deregibus MC, Bruno S, Grange C, Fonsato V, Tetta C. Exosome/microvesicle-mediated epigenetic reprogramming of cells. Am J Cancer Res 2011;1:98–110. PMID: 21969178.

15. Rak J, Guha A. Extracellular vesicles: vehicles that spread cancer genes. Bioessays 2012;34:489–497. PMID: 22442051.


16. Cai Z, Yang F, Yu L, Yu Z, Jiang L, Wang Q, et al. Activated T cell exosomes promote tumor invasion via Fas signaling pathway. J Immunol 2012;188:5954–5961. PMID: 22573809.


17. Bellingham SA, Guo BB, Coleman BM, Hill AF. Exosomes: vehicles for the transfer of toxic proteins associated with neurodegenerative diseases? Front Physiol 2012;3:124. PMID: 22563321.



18. Emmanouilidou E, Melachroinou K, Roumeliotis T, Garbis SD, Ntzouni M, Margaritis LH, et al. Cell-produced alpha-synuclein is secreted in a calcium-dependent manner by exosomes and impacts neuronal survival. J Neurosci 2010;30:6838–6851. PMID: 20484626.



19. Aswad H, Forterre A, Wiklander OP, Vial G, Danty-Berger E, Jalabert A, et al. Exosomes participate in the alteration of muscle homeostasis during lipid-induced insulin resistance in mice. Diabetologia 2014;57:2155–2164. PMID: 25073444.



20. Jayachandran M, Litwiller RD, Lahr BD, Bailey KR, Owen WG, Mulvagh SL, et al. Alterations in platelet function and cell-derived microvesicles in recently menopausal women: relationship to metabolic syndrome and atherogenic risk. J Cardiovasc Transl Res 2011;4:811–822. PMID: 21786187.



21. Diamant M, Nieuwland R, Pablo RF, Sturk A, Smit JW, Radder JK. Elevated numbers of tissue-factor exposing microparticles correlate with components of the metabolic syndrome in uncomplicated type 2 diabetes mellitus. Circulation 2002;106:2442–2447. PMID: 12417540.


22. D'Alessandra Y, Devanna P, Limana F, Straino S, Di Carlo A, Brambilla PG, et al. Circulating microRNAs are new and sensitive biomarkers of myocardial infarction. Eur Heart J 2010;31:2765–2773. PMID: 20534597.




23. Feng B, Chen Y, Luo Y, Chen M, Li X, Ni Y. Circulating level of microparticles and their correlation with arterial elasticity and endothelium-dependent dilation in patients with type 2 diabetes mellitus. Atherosclerosis 2010;208:264–269. PMID: 19674745.


24. Kim SJ, Moon GJ, Cho YH, Kang HY, Hyung NK, Kim D, et al. Circulating mesenchymal stem cells microparticles in patients with cerebrovascular disease. PLoS One 2012;7:e37036. PMID: 22615882.



25. Ogata M, Inanami O, Nakajima M, Nakajima T, Hiraoka W, Kuwabara M. Ca(2+)-dependent and caspase-3-independent apoptosis caused by damage in Golgi apparatus due to 2,4,5,7-tetrabromorhodamine 123 bromide-induced photodynamic effects. Photochem Photobiol 2003;78:241–247. PMID: 14556310.


26. Safdar A, Saleem A, Tarnopolsky MA. The potential of endurance exercise-derived exosomes to treat metabolic diseases. Nat Rev Endocrinol 2016;12:504–517. PMID: 27230949.


27. Jalabert A, Vial G, Guay C, Wiklander OP, Nordin JZ, Aswad H, et al. Exosome-like vesicles released from lipid-induced insulin-resistant muscles modulate gene expression and proliferation of beta recipient cells in mice. Diabetologia 2016;59:1049–1058. PMID: 26852333.



28. Lee JE, Moon PG, Lee IK, Baek MC. Proteomic analysis of extracellular vesicles released by adipocytes of Otsuka Long-Evans Tokushima Fatty (OLETF) Rats. Protein J 2015;34:220–235. PMID: 25998041.


29. Santovito D, De Nardis V, Marcantonio P, Mandolini C, Paganelli C, Vitale E, et al. Plasma exosome microRNA profiling unravels a new potential modulator of adiponectin pathway in diabetes: effect of glycemic control. J Clin Endocrinol Metab 2014;99:E1681–E1685. PMID: 24937531.



30. Deng ZB, Poliakov A, Hardy RW, Clements R, Liu C, Liu Y, et al. Adipose tissue exosome-like vesicles mediate activation of macrophage-induced insulin resistance. Diabetes 2009;58:2498–2505. PMID: 19675137.



31. Jansen F, Yang X, Hoelscher M, Cattelan A, Schmitz T, Proebsting S, et al. Endothelial microparticle-mediated transfer of MicroRNA-126 promotes vascular endothelial cell repair via SPRED1 and is abrogated in glucose-damaged endothelial microparticles. Circulation 2013;128:2026–2038. PMID: 24014835.


32. Wang X, Huang W, Liu G, Cai W, Millard RW, Wang Y, et al. Cardiomyocytes mediate anti-angiogenesis in type 2 diabetic rats through the exosomal transfer of miR-320 into endothelial cells. J Mol Cell Cardiol 2014;74:139–150. PMID: 24825548.



33. O'Neill S, Bohl M, Gregersen S, Hermansen K, O'Driscoll L. Blood-based biomarkers for metabolic syndrome. Trends Endocrinol Metab 2016;27:363–374. PMID: 27150849.


34. Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin Pharmacol Ther 2001;69:89–95. PMID: 11240971.


35. Lawson C, Vicencio JM, Yellon DM, Davidson SM. Microvesicles and exosomes: new players in metabolic and cardiovascular disease. J Endocrinol 2016;228:R57–R71. PMID: 26743452.


36. Koeck ES, Iordanskaia T, Sevilla S, Ferrante SC, Hubal MJ, Freishtat RJ, et al. Adipocyte exosomes induce transforming growth factor beta pathway dysregulation in hepatocytes: a novel paradigm for obesity-related liver disease. J Surg Res 2014;192:268–275. PMID: 25086727.


37. Nicklas BJ, Penninx BW, Cesari M, Kritchevsky SB, Newman AB, Kanaya AM, et al. Association of visceral adipose tissue with incident myocardial infarction in older men and women: the Health, Aging and Body Composition Study. Am J Epidemiol 2004;160:741–749. PMID: 15466496.



38. Kanhai DA, Kappelle LJ, van der Graaf Y, Uiterwaal CS, Visseren FL. SMART Study Group. The risk of general and abdominal adiposity in the occurrence of new vascular events and mortality in patients with various manifestations of vascular disease. Int J Obes (Lond) 2012;36:695–702. PMID: 21712810.


39. Livshits MA, Khomyakova E, Evtushenko EG, Lazarev VN, Kulemin NA, Semina SE, et al. Corrigendum: Isolation of exosomes by differential centrifugation: Theoretical analysis of a commonly used protocol. Sci Rep 2016;6:21447. PMID: 27022711.




40. Cocucci E, Racchetti G, Podini P, Meldolesi J. Enlargeosome traffic: exocytosis triggered by various signals is followed by endocytosis, membrane shedding or both. Traffic 2007;8:742–757. PMID: 17488290.


41. Eken C, Gasser O, Zenhaeusern G, Oehri I, Hess C, Schifferli JA. Polymorphonuclear neutrophil-derived ectosomes interfere with the maturation of monocyte-derived dendritic cells. J Immunol 2008;180:817–824. PMID: 18178820.


42. Lötvall J, Hill AF, Hochberg F, Buzás EI, Di Vizio D, Gardiner C, et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J Extracell Vesicles 2014;3:26913. PMID: 25536934.



43. Linares R, Tan S, Gounou C, Arraud N, Brisson AR. High-speed centrifugation induces aggregation of extracellular vesicles. J Extracell Vesicles 2015;4:29509. PMID: 26700615.



44. Van Deun J, Mestdagh P, Sormunen R, Cocquyt V, Vermaelen K, Vandesompele J, et al. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J Extracell Vesicles 2014;3:24858.



45. Kalra H, Adda CG, Liem M, Ang CS, Mechler A, Simpson RJ, et al. Comparative proteomics evaluation of plasma exosome isolation techniques and assessment of the stability of exosomes in normal human blood plasma. Proteomics 2013;13:3354–3364. PMID: 24115447.


46. Hong CS, Funk S, Muller L, Boyiadzis M, Whiteside TL. Isolation of biologically active and morphologically intact exosomes from plasma of patients with cancer. J Extracell Vesicles 2016;5:29289. PMID: 27018366.



47. Fiandaca MS, Kapogiannis D, Mapstone M, Boxer A, Eitan E, Schwartz JB, et al. Identification of preclinical Alzheimer's disease by a profile of pathogenic proteins in neurally derived blood exosomes: A case-control study. Alzheimers Dement 2015;11:600–607.e1. PMID: 25130657.



48. Shi M, Liu C, Cook TJ, Bullock KM, Zhao Y, Ginghina C, et al. Plasma exosomal α-synuclein is likely CNS-derived and increased in Parkinson's disease. Acta Neuropathol 2014;128:639–650. PMID: 24997849.



49. Lobb RJ, Becker M, Wen SW, Wong CS, Wiegmans AP, Leimgruber A, et al. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J Extracell Vesicles 2015;4:27031. PMID: 26194179.



50. Agouni A, Ducluzeau PH, Benameur T, Faure S, Sladkova M, Duluc L, et al. Microparticles from patients with metabolic syndrome induce vascular hypo-reactivity via Fas/Fasligand pathway in mice. PLoS One 2011;6:e27809. PMID: 22110764.



51. Agouni A, Lagrue-Lak-Hal AH, Ducluzeau PH, Mostefai HA, Draunet-Busson C, Leftheriotis G, et al. Endothelial dysfunction caused by circulating microparticles from patients with metabolic syndrome. Am J Pathol 2008;173:1210–1219. PMID: 18772329.



52. Amabile N, Cheng S, Renard JM, Larson MG, Ghorbani A, McCabe E, et al. Association of circulating endothelial microparticles with cardiometabolic risk factors in the Framingham Heart Study. Eur Heart J 2014;35:2972–2979. PMID: 24742886.




53. Chen Y, Feng B, Li X, Ni Y, Luo Y. Plasma endothelial microparticles and their correlation with the presence of hypertension and arterial stiffness in patients with type 2 diabetes. J Clin Hypertens (Greenwich) 2012;14:455–460. PMID: 22747618.



54. Kranendonk ME, de Kleijn DP, Kalkhoven E, Kanhai DA, Uiterwaal CS, van der Graaf Y, et al. Extracellular vesicle markers in relation to obesity and metabolic complications in patients with manifest cardiovascular disease. Cardiovasc Diabetol 2014;13:37. PMID: 24498934.



55. Burger D, Thibodeau JF, Holterman CE, Burns KD, Touyz RM, Kennedy CR. Urinary podocyte microparticles identify prealbuminuric diabetic glomerular injury. J Am Soc Nephrol 2014;25:1401–1407. PMID: 24676640.



56. Sun AL, Deng JT, Guan GJ, Chen SH, Liu YT, Cheng J, et al. Dipeptidyl peptidase-IV is a potential molecular biomarker in diabetic kidney disease. Diab Vasc Dis Res 2012;9:301–308. PMID: 22388283.


57. Zubiri I, Posada-Ayala M, Sanz-Maroto A, Calvo E, Martin-Lorenzo M, Gonzalez-Calero L, et al. Diabetic nephropathy induces changes in the proteome of human urinary exosomes as revealed by label-free comparative analysis. J Proteomics 2014;96:92–102. PMID: 24211404.


Fig. 1
Biogenesis and secretion of exosomes. Exosomes originate from the endolysosomal pathway and exhibit a homogeneous size and shape. During the maturation of the early endosome, the inward budding of cellular compartments that occurs in the membrane of the endosome produces multivesicular bodies (MVBs). The internal vesicles (intraluminal vesicles; ILVs) within MVBs contain soluble proteins, transmembrane proteins, lipids, receptors, ligands, mRNAs and miRNAs lower inserted box that are responsible for intercellular communication. The internal sorting of these molecules is precisely regulated by several mechanisms including the endosomal sorting complexes required for transport system. MVBs can fuse with lysosomes to recycle the contents of the ILVs or with the plasma membrane to release ILVs as exosomes. The mechanisms that decide between the release of MVBs as exosomes or their allocation to recycling pathways are not fully understood. GPI, glycophosphatidylinositol; TM, transmembrane.
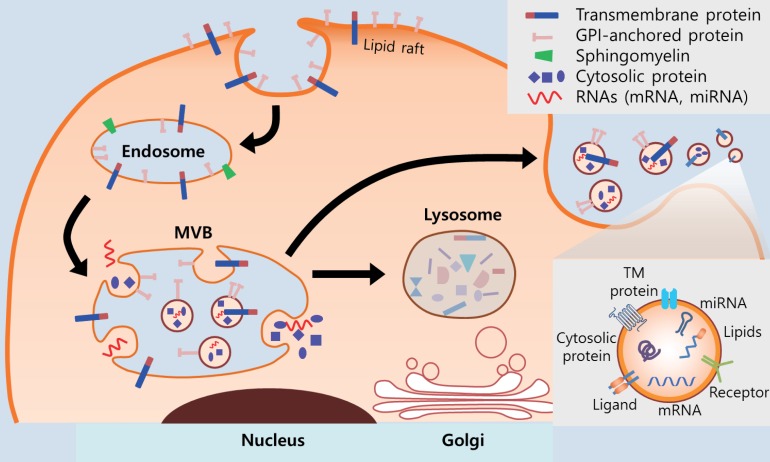
Fig. 2
Electron microscopic image of exosomes prepared from cell culture media (CM). After full differentiation of murine myoblast (C2C12 cells) to myocytes, exosomes were isolated from CM by differential ultracentrifugation. (A, B) Small vesicles (white arrows) homogeneously sized <150 nm are clearly identifiable. (C, D) Several aggregated vesicles are also observed when ultracentrifugation is used to prepare exosomes. Arrowheads and asterisk indicate membrane of exosome and exosomal body, respectively.

- TOOLS
- Related articles in APEM






